Scale Up Bulk Similarity Calculations for Sparse Embeddings
Last Updated on April 21, 2023 by Editorial Team
Author(s): Rodrigo Agundez
Originally published on Towards AI.
ChunkDot support for sparse matrices
In my previous blog post, I introduced ChunkDot, a library that performs multi-threaded matrix multiplication and cosine similarity. ChunkDot is appropriate for calculating the K most similar items for a large number of items by chunking the item matrix representation (embeddings) and using Numba to accelerate the calculations.
Cosine Similarity for 1 Trillion Pairs of Vectors
Introducing ChunkDot
pub.towardsai.net
I described how ChunkDot works under the hood and I showed some benchmarks against execution time and memory consumption for dense embeddings. I recently extended the support to sparse matrices, this blog post aims to explain the methodology and show an example of how you could use it.
If you would like to skip the explanation and jump directly into the usage instructions:
chunkdot
Multi-threaded matrix multiplication and cosine similarity calculations. Appropriate for the calculation of the K most…
pypi.org
Motivation
The calculation of cosine similarity using sparse vector representations suffers from the same issue as using dense vector representations. Even though the items are represented as a sparse embedding matrix, all pairwise similarity calculations might be non-zero. If you have N items, the calculation of pairwise similarities will yield a dense similarity matrix N x N, with a memory footprint that scales as N². For 100K items, this is ~80GB, and for 1 million ~8000GB.
ChunkDot splits the embeddings matrix in chunks and parallelizes the calculation of cosine similarity, retrieving the K most similar items per item. The diagram below shows how ChunkDot works, but please refer to my previous blog post for details.
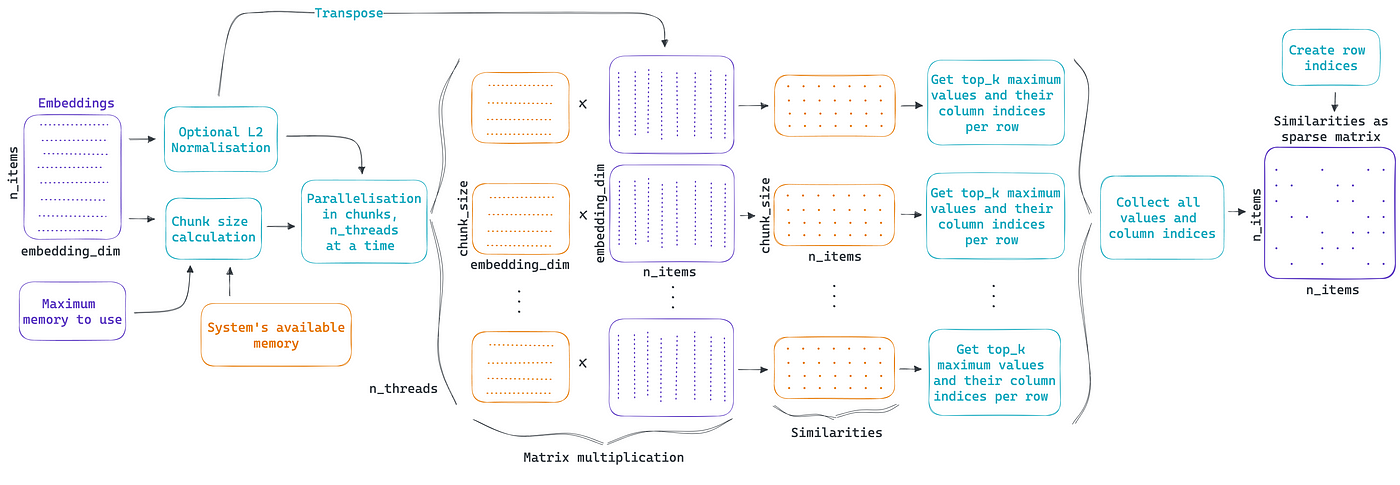
There are several use cases that perform similarity calculations over sparse vector representations, for example, it often happens in NLP that the vectorization of a corpus of text is done via the use of tokens, n-grams, or words. This results in a highly sparse matrix representation where the vocabulary for the total corpus is much bigger than the vocabulary for each document. By extending the support of ChunkDot to sparse matrices, ChunkDot can help scale up these calculations and support uses cases like:
- Deduplication of documents.
- Recommendations for similar documents.
- Clustering / Tagging / Grouping of documents.
ChunkDot for Sparse Matrices
To support sparse matrices, ChunkDot uses the same Numba multi-threading methodology. The new release adds a chunked sparse matrix multiplication logic compatible with Numba in no-python mode. Then this sparse matrix multiplication logic is embedded in each parallel thread to calculate the chunked cosine similarity.

I wanted to use SciPy’s sparse matrix multiplication implementation but unfortunately, SciPy is not supported by Numba, so I had to write the logic myself. I made the decision to support all SciPy sparse matrix formats but, under the hood, perform the matrix multiplication in the CSR format; it looks something like this:
values = np.zeros((left_n_rows, right_n_cols))
for row_left in range(left_n_rows):
for left_i in range(matrix_left_indptr[row_left], matrix_left_indptr[row_left + 1]):
col_left = matrix_left_indices[left_i]
value_left = matrix_left_data[left_i]
for right_i in range(matrix_right_indptr[col_left], matrix_right_indptr[col_left + 1]):
col_right = matrix_right_indices[right_i]
value_right = matrix_right_data[right_i]
values[row_left, col_right] += value_left * value_right
I you are not familiar with sparse matrices, the logic can seem a bit challenging to understand. It is, at least to me. In short, what the logic above does is to get, per row, the column index of each non-zero item in the left matrix and then jump directly into the row of the right matrix for that column index, finally aggregate to the matrix entry the multiplication of both values.
Cosine Similarity over a blog post corpus
In this example, I concentrate only on the application of ChunkDot for similarity calculations. I do not put any effort into cleaning or normalizing the corpus as it should be done in an actual use case.
Let’s read 100K blog posts and take a look at some examples:
import pandas as pd
blogs = pd.read_csv("blogtext.csv", usecols=["text"], nrows=100000)
for _, text in blogs.sample(3).iterrows():
print(text.text, "\n")
“Cordy has her moments. You know what owns? ‘Inside Out’ by urlLink Eve 6 . I’m going to really make myself look young here and just say that song is one of the anthems of my generation. I rocked out to it to the point of nearly losing my voice on the way home from work today. It kicks that much ass. You know what else owns? A good tuna melt. I wish I could be happy today not feeling so guilty about how badly we’re getting our asses beat right now. “
“On Bubb Rubb and Woo Woo….. To fully grasp the wonderful bit of comedy that is Bubb Rubb and Woo Woo, you must first watch a little urlLink News Clip from Oakland. After you watch the clip, go directly to the urlLink Bubb Rubb site and mix it up for yourself…..let me know what you think…I wasted like 20 minutes playing around with it….enjoy! “
“This is What I heard today (meaning he said it for the first time AFAIK) > pizza (pee-zah)(pee-tah) sheep (seep) kick wagon paper printer (pin-tah) fishie movie sit down (sih down) bounce stick — — — — — — I think that’s it. He’s repeating so many things lately, and coming out with so many words I’ve never heard him use that I just can’t keep track anymore. It’s unbelievable how his vocabulary, or at least his attempt at using his existing vocabulary, has just exploded lately. He’s going to be a talking machine in no time.”
The blog posts are rich in content and in the number of words used, a good realistic corpus for our example. The plot below shows that this dataset has texts that spread across the spectrum in terms of the number of words used.
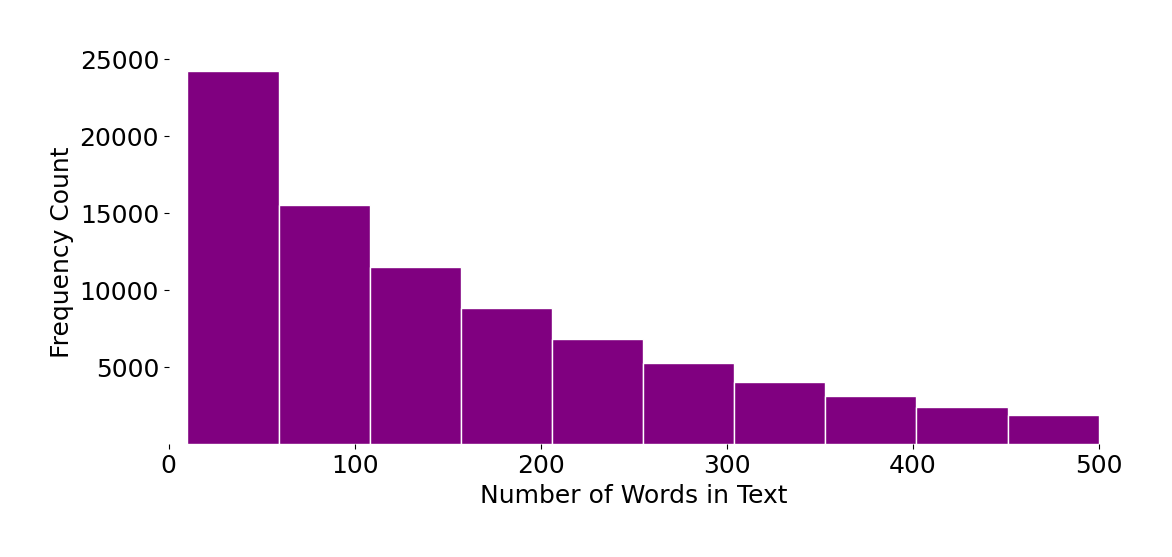
Let’s now vectorize the corpus. Here I use SciKit-Learn’s TfidfVectorizer for simplicity but this logic can be substituted with anything that will yield a sparse matrix representation of the corpus of text.
from sklearn.feature_extraction.text import TfidfVectorizer
vectorizer = TfidfVectorizer(analyzer="word", stop_words="english")
embeddings = vectorizer.fit_transform(blogs["text"])
embeddings
<100000x214146 sparse matrix of type '<class 'numpy.float64'>'
with 6817666 stored elements in Compressed Sparse Row format>
The resulting embeddings matrix consists of 100K rows representing each blog post and 214K columns representing each word in the corpus vocabulary. If the embeddings were represented in a dense form, they would occupy ~171GB of memory, instead the embeddings are represented as a sparse matrix with a density of ~0.003.
Let’s now calculate the 10 most similar blog posts per blog post using ChunkDot.
from chunkdot import cosine_similarity_top_k
similarities = cosine_similarity_top_k(embeddings, top_k=10)
similarities
<100000x100000 sparse matrix of type '<class 'numpy.float64'>'
with 1000000 stored elements in Compressed Sparse Row format>
The cosine similarity calculations take ~1min.
from timeit import timeit
timeit(lambda: cosine_similarity_top_k(embeddings, top_k=10), number=1)
67.77648650599997
To sparse or not to sparse, that is the question
Since embeddings can be represented as dense or sparse and the matrix multiplication logic is different for both, is natural to ask, should I always use a sparse representation? The answer is no.
There is a performance penalty on calculations over sparse matrices that are not sparse enough. Below is a comparison of matrix multiplication performance when using a dense matrix and a dense matrix multiplication versus when using a sparse matrix and a sparse matrix multiplication. The results are presented as a ratio, a value bigger than one means that the sparse representation is faster.

As we can see, using sparse representations make sense only for densities below ~0.03. For higher densities, the benefits are quickly reduced, but at the same time, for big matrices and low densities ~.001, there is a 100x benefit of using a sparse representation. In the NLP example above, the embedding matrix density was ~0.003.
Conclusion
I hope you find this blog post and ChunkDot useful! I enjoyed adding the sparse matrix support. Some potential improvements:
- Extend the functionality to 2 different input matrices. Calculate the cosine similarity between the rows of a matrix and the rows of another one.
- Instead of returning the K most similar items, return items whose similarity is above/below a certain threshold.
- Add GPU support since Numba supports it.
Numba for CUDA GPUs – Numba 0.56.4+0.g288a38bbd.dirty-py3.7-linux-x86_64.egg documentation
Callback into the Python Interpreter from within JIT'ed code
numba.readthedocs.io
Join thousands of data leaders on the AI newsletter. Join over 80,000 subscribers and keep up to date with the latest developments in AI. From research to projects and ideas. If you are building an AI startup, an AI-related product, or a service, we invite you to consider becoming a sponsor.
Published via Towards AI
Take our 90+ lesson From Beginner to Advanced LLM Developer Certification: From choosing a project to deploying a working product this is the most comprehensive and practical LLM course out there!
Towards AI has published Building LLMs for Production—our 470+ page guide to mastering LLMs with practical projects and expert insights!

Discover Your Dream AI Career at Towards AI Jobs
Towards AI has built a jobs board tailored specifically to Machine Learning and Data Science Jobs and Skills. Our software searches for live AI jobs each hour, labels and categorises them and makes them easily searchable. Explore over 40,000 live jobs today with Towards AI Jobs!
Note: Content contains the views of the contributing authors and not Towards AI.















