Data Exploration with Python: A Hands-On Demo in EDA (and Why It’s Essential for Model Building)
Last Updated on December 2, 2025 by Editorial Team
Author(s): Faizulkhan
Originally published on Towards AI.
Exploratory Data Analysis (EDA) is the bridge between raw data and reliable machine-learning models. In this post, we will learn the “why” and the “how” through a complete, runnable example in Python using NumPy, Pandas, Matplotlib, SciPy, and Scikit-Learn. We’ll detect and fix missing values, spot outliers, visualize distributions, compare variables, normalize features, quantify correlations, and fit a simple regression — then turn those insights into a tiny predictive function.
What We will Learn
- Why EDA is the critical first step before modeling.
- The difference between Python lists and NumPy arrays (and why arrays are better for math).
- How to use Pandas DataFrames for loading, cleaning, filtering, and grouping data.
- How to visualize distributions (histograms, boxplots, density plots).
- Measures of central tendency (mean/median/mode) and variance (range/variance/std).
- Outlier detection and removal with quantiles.
- Feature normalization (Min-Max scaling) and correlation analysis.
- Simple linear regression (least squares), interpretation, and a tiny prediction function.
- A practical workflow you can replicate for new datasets.
The Dataset
We’ll use a small “student grades” dataset with two numeric variables:
StudyHours— average weekly study timeGrade— final course grade
We can fetch it with:
curl -o data/grades.csv "https://raw.githubusercontent.com/faizulkhan56/python-eda-discovery/main/grades.csv"
# or
wget -O data/grades.csv "https://raw.githubusercontent.com/faizulkhan56/python-eda-discovery/main/grades.csv"
(Or run the included scripts/download_data.py if you’re using the linked project.)
Quick Environment Setup (One Time)
Create a project folder and a virtual environment:
python3 -m venv .venv
source .venv/bin/activate # Windows: .\.venv\Scripts\Activate.ps1
pip install --upgrade pip
Install dependencies:
pip install jupyter ipykernel numpy pandas matplotlib scipy scikit-learn
python -m ipykernel install --user --name python-eda-student-grades --display-name "Python (EDA: Student Grades)"
Open Jupyter and select the kernel “Python (EDA: Student Grades)”:
jupyter notebook

Why EDA Matters (Before We Train a Model)
EDA is how we learn the story of our data is trying to tell. Before fitting models, We need to answer:
- What’s missing or broken?
Missing values, strange encodings, inconsistent units, duplicated rows. - What’s the distribution?
Are variables normal, skewed, multimodal? This impacts transformations, model choice, and evaluation. - Where are the outliers?
Anomalies can warp means, inflate variance, and mislead models. - What relates to what?
Correlations and relationships suggest useful features (and reveal leakage risks). - Are scales compatible?
Many algorithms are sensitive to feature scaling. Normalization or standardization may be required. - What’s the hypothesis?
EDA shapes the modeling plan: which features to engineer, which algorithms to try, and what to watch out for.
Outcome: Cleaner data, better features, realistic expectations, and fewer model “mysteries.”
Step 1 — NumPy vs Python Lists
Start with raw Python data:
data = [50,50,47,97,49,3,53,42,26,74,82,62,37,15,70,27,36,35,48,52,63,64]
print(data)
import numpy as np
grades = np.array(data)
print(type(data), 'x 2:', data * 2) # list repeats
print(type(grades), 'x 2:', grades * 2) # array multiplies element-wise
Key idea: Lists are general-purpose; NumPy arrays are optimized for math. Arrays enable vectorized operations (fast, clean, bug-resistant).

Step 2 — From Arrays to Tables with Pandas
Create a tabular structure that’s ideal for analysis:
import numpy as np
import pandas as pd
study_hours = [10.0,11.5,9.0,16.0,9.25,1.0,11.5,9.0,8.5,14.5,15.5,
13.75,9.0,8.0,15.5,8.0,9.0,6.0,10.0,12.0,12.5,12.0]
grades = np.array(data)
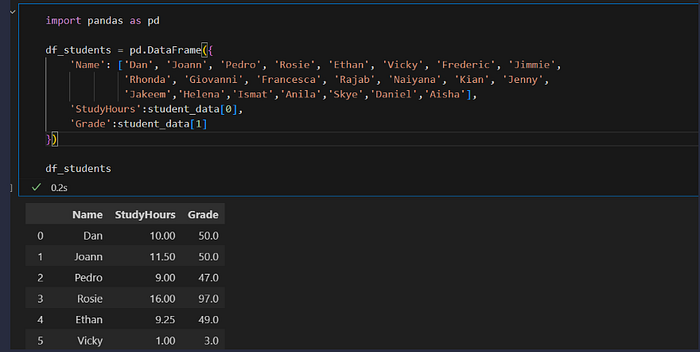
df_students = pd.DataFrame({
'Name': ['Dan','Joann','Pedro','Rosie','Ethan','Vicky','Frederic','Jimmie',
'Rhonda','Giovanni','Francesca','Rajab','Naiyana','Kian','Jenny',
'Jakeem','Helena','Ismat','Anila','Skye','Daniel','Aisha'],
'StudyHours': study_hours,
'Grade': grades
})
df_students.head()
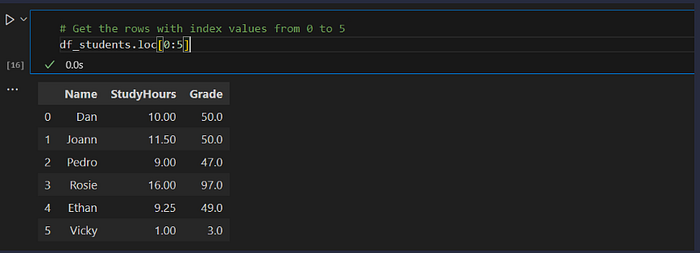
Loc vs Iloc:
df_students.loc[0:5] # by index *label* (inclusive end)
df_students.iloc[0:5] # by row *position* (exclusive end)
df_students.iloc[0, [1,2]] # pick by row/column position
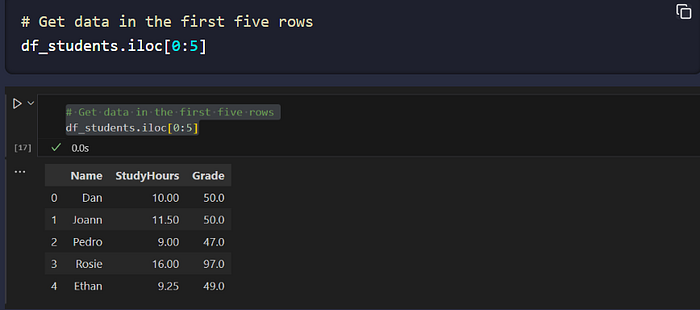
Can We notice the difference between the two methods?
- The loc method returned rows with index label in the list of values from 0 to 5 — which includes 0, 1, 2, 3, 4, and 5 (six rows). However, the iloc method returns the rows in the positions included in the range 0 to 5, and since integer ranges don’t include the upper-bound value, this includes positions 0, 1, 2, 3, and 4 (five rows).
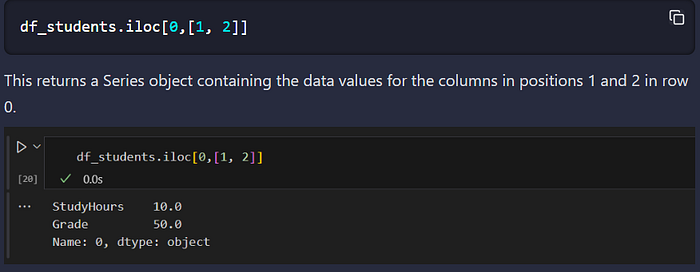
- iloc identifies data values in a DataFrame by position, which extends beyond rows to columns. So for example, you can use it to find the values for the columns in positions 1 and 2 in row 0, like this:
Step 3 — Load from CSV and Handle Missing Values
In real life, we will read from files and face missingness:
df_students = pd.read_csv('grades.csv', delimiter=',', header=0)
# Detect missing values
df_students.isnull().sum()
# See the actual rows with any nulls
df_students[df_students.isnull().any(axis=1)]
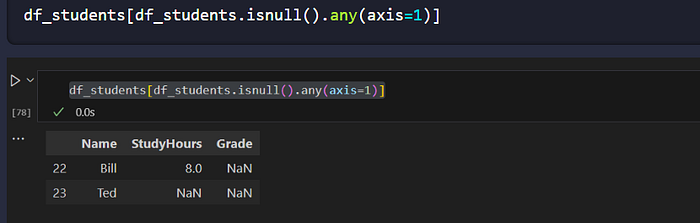
Two common strategies:
- Impute (e.g., fill with the mean for continuous variables)
- Drop rows (when safe and justified)
df_students.StudyHours = df_students.StudyHours.fillna(df_students.StudyHours.mean())
df_students = df_students.dropna(axis=0, how='any')
axis=0 means that the rows are dropped, and how='any' means that any rows with missing values will be dropped.
Step 4 — Explore Central Tendency & Spread
Central tendency: mean, median, mode
Spread: range, variance, standard deviation
var = df_students['Grade']
stats = {
"min": var.min(),
"mean": var.mean(),
"median": var.median(),
"mode": var.mode()[0],
"max": var.max()
}
stats
Visualize distributions:
from matplotlib import pyplot as plt
%matplotlib inline
plt.figure(figsize=(10,4))
plt.hist(df_students['Grade'])
plt.title('Grade Distribution'); plt.xlabel('Value'); plt.ylabel('Frequency')
plt.show()
plt.figure(figsize=(10,4))
plt.boxplot(df_students['Grade'])
plt.title('Grade Boxplot'); plt.show()
Why this matters for modeling:
- If features are normal-ish, linear models may be appropriate.
- Skewed features might benefit from transformations.
- Boxplots and histograms quickly reveal issues to address before training.
Step 5 — Outliers and Robustness
Spotting outliers via quantiles:
In most real-world cases, it’s easier to consider outliers as being values that fall below or above percentiles within which most of the data lie. For example, the following code uses the Pandas quantile function to exclude observations below the 0.01th percentile (the value above which 99% of the data reside).
q01 = df_students.StudyHours.quantile(0.01)
df_no_low_outliers = df_students[df_students.StudyHours > q01]
With the outliers removed, the box plot shows all data within the four quartiles. Note that the distribution is not symmetric like it is for the grade data though — there are some students with very high study times of around 16 hours, but the bulk of the data is between 7 and 13 hours; The few extremely high values pull the mean towards the higher end of the scale.

Re-plot to verify improvements.
Impact on models: Outliers can dominate loss functions (e.g., squared errors), skew learned parameters, and hurt generalization.
Measure Of variance
So now we have a good idea where the middle of the grade and study hours data distributions are. However, there’s another aspect of the distributions we should examine: how much variability is there in the data?
Typical statistics that measure variability in the data include:
- Range: The difference between the maximum and minimum. There’s no built-in function for this, but it’s easy to calculate using the min and max functions.
- Variance: The average of the squared difference from the mean. You can use the built-in var function to find this.
- Standard Deviation: The square root of the variance. You can use the built-in std function to find this.
Typical statistics that measure variability in the data include:
- Range: The difference between the maximum and minimum. There’s no built-in function for this, but it’s easy to calculate using the min and max functions.
- Variance: The average of the squared difference from the mean. You can use the built-in var function to find this.
- Standard Deviation: The square root of the variance. You can use the built-in std function to find this.
for col_name in ['Grade','StudyHours']:
col = df_students[col_name]
rng = col.max() - col.min()
var = col.var()
std = col.std()
print('\n{}:\n - Range: {:.2f}\n - Variance: {:.2f}\n - Std.Dev: {:.2f}'.format(col_name, rng, var, std))
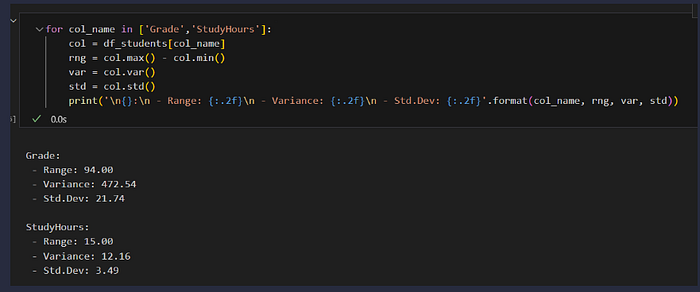
Of these statistics, the standard deviation is generally the most useful. It provides a measure of variance in the data on the same scale as the data itself (so grade points for the Grade distribution and hours for the StudyHours distribution). The higher the standard deviation, the more variance there is when comparing values in the distribution to the distribution mean — in other words, the data is more spread out.
When working with a normal distribution, the standard deviation works with the particular characteristics of a normal distribution to provide even greater insight. Run the cell below to see the relationship between standard deviations and the data in the normal distribution.
import scipy.stats as stats
# Get the Grade column
col = df_students['Grade']
# get the density
density = stats.gaussian_kde(col)
# Plot the density
col.plot.density()
# Get the mean and standard deviation
s = col.std()
m = col.mean()
# Annotate 1 stdev
x1 = [m-s, m+s]
y1 = density(x1)
plt.plot(x1,y1, color='magenta')
plt.annotate('1 std (68.26%)', (x1[1],y1[1]))
# Annotate 2 stdevs
x2 = [m-(s*2), m+(s*2)]
y2 = density(x2)
plt.plot(x2,y2, color='green')
plt.annotate('2 std (95.45%)', (x2[1],y2[1]))
# Annotate 3 stdevs
x3 = [m-(s*3), m+(s*3)]
y3 = density(x3)
plt.plot(x3,y3, color='orange')
plt.annotate('3 std (99.73%)', (x3[1],y3[1]))
# Show the location of the mean
plt.axvline(col.mean(), color='cyan', linestyle='dashed', linewidth=1)
plt.axis('off')
plt.show()
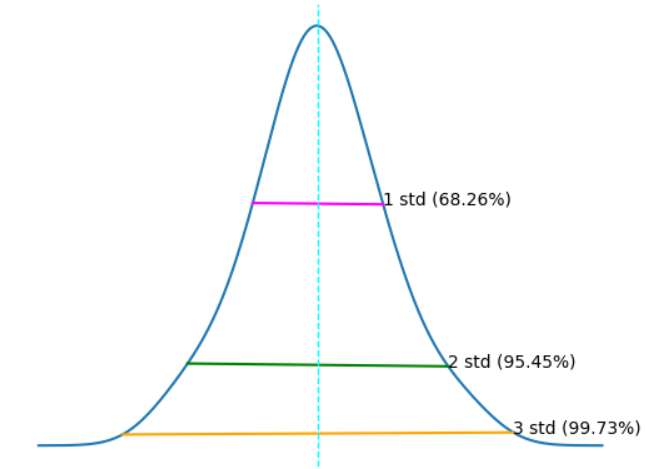
The horizontal lines show the percentage of data within 1, 2, and 3 standard deviations of the mean (plus or minus).
In any normal distribution:
- Approximately 68.26% of values fall within one standard deviation from the mean.
- Approximately 95.45% of values fall within two standard deviations from the mean.
- Approximately 99.73% of values fall within three standard deviations from the mean.
So, since we know that the mean grade is 49.18, the standard deviation is 21.74, and distribution of grades is approximately normal; we can calculate that 68.26% of students should achieve a grade between 27.44 and 70.92.
The descriptive statistics we’ve used to understand the distribution of the student data variables are the basis of statistical analysis; and because they’re such an important part of exploring your data, there’s a built-in Describe method of the DataFrame object that returns the main descriptive statistics for all numeric columns.
Variance — Formula and Intuition
Variance (σ²) measures how far the data points are spread out from the mean (average).
Mathematically, for a dataset of nnn values x1,x2,x3,…,xnx_1, x_2, x_3, …, x_nx1,x2,x3,…,xn:
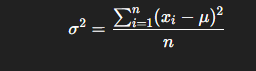
where
- xi = each data point
- μ = mean of all data points
- n= number of observations
👉 This gives the average squared deviation from the mean.
for samples
When we’re working with a sample (not the full population), we divide by n−1 instead of n to make the estimate unbiased:
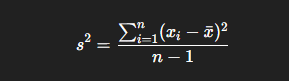
STD deviation
Standard deviation (σ) is simply the square root of variance:
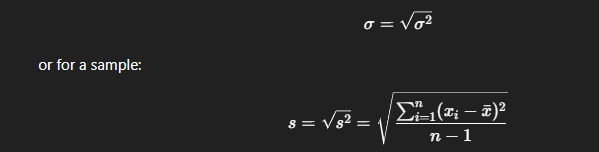
Step 6 — Grouping, Aggregation, and New Columns
Add a Pass/Fail flag and explore:
passes = pd.Series(df_students['Grade'] >= 60)
df_students = pd.concat([df_students, passes.rename('Pass')], axis=1)
# Counts and group means
df_students.groupby('Pass').Name.count()
df_students.groupby('Pass')[['StudyHours','Grade']].mean()
Quick bar and pie charts help stakeholders grasp outcomes immediately.
we can use the pyplot class from Matplotlib to plot the chart. This class provides a whole bunch of ways to improve the visual elements of the plot. For example, the following code:
- Specifies the color of the bar chart.
- Adds a title to the chart (so we know what it represents)
- Adds labels to the X and Y (so we know which axis shows which data)
- Adds a grid (to make it easier to determine the values for the bars)
- Rotates the X markers (so we can read them)
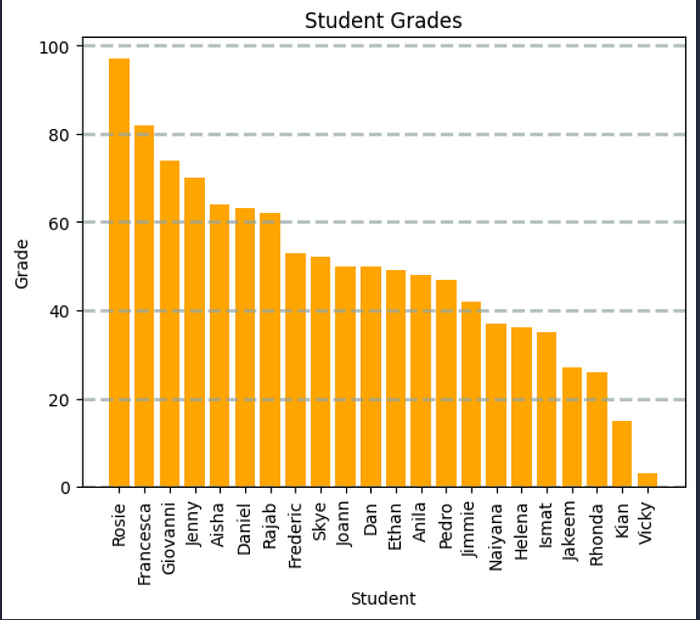
# Create a bar plot of name vs grade
plt.bar(x=df_students.Name, height=df_students.Grade, color='orange')
# Customize the chart
plt.title('Student Grades')
plt.xlabel('Student')
plt.ylabel('Grade')
plt.grid(color='#95a5a6', linestyle='--', linewidth=2, axis='y', alpha=0.7)
plt.xticks(rotation=90)
# Display the plot
plt.show()
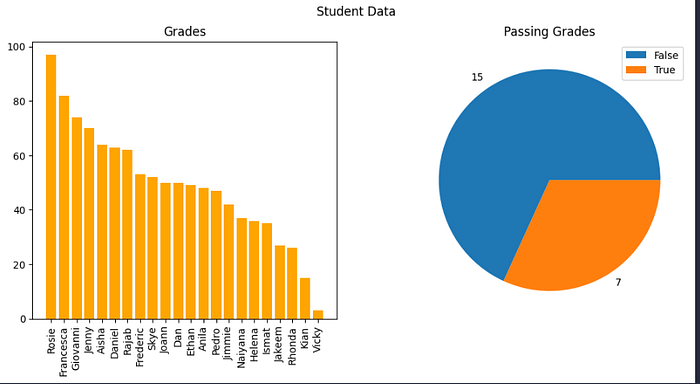
# Create a figure for 2 subplots (1 row, 2 columns)
fig, ax = plt.subplots(1, 2, figsize = (12,5))
# Create a bar plot of name vs grade on the first axis
ax[0].bar(x=df_students.Name, height=df_students.Grade, color='orange')
ax[0].set_title('Grades')
ax[0].set_xticklabels(df_students.Name, rotation=90)
# Create a pie chart of pass counts on the second axis
pass_counts = df_students['Pass'].value_counts()
ax[1].pie(pass_counts, labels=pass_counts)
ax[1].set_title('Passing Grades')
ax[1].legend(pass_counts.keys().tolist())
# Add a title to the Figure
fig.suptitle('Student Data')
# Show the figure
fig.show()
Step 7 — Compare Variables & Normalize Scales
Plot Grade vs StudyHours side-by-side:
df_students.plot(x='Name', y=['Grade','StudyHours'], kind='bar', figsize=(8,5))
Because they’re on different scales, use Min-Max scaling:
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler()
df_norm = df_students[['Name','Grade','StudyHours']].copy()
df_norm[['Grade','StudyHours']] = scaler.fit_transform(df_norm[['Grade','StudyHours']])
df_norm.plot(x='Name', y=['Grade','StudyHours'], kind='bar', figsize=(8,5))
Why scale? Many algorithms (KNN, SVM, gradient descent dynamics) are sensitive to feature magnitude. Scaling reduces numerical instability and training surprises.
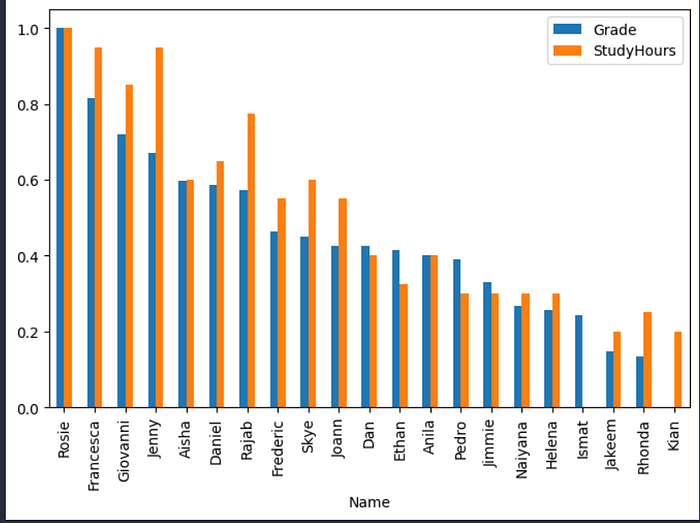
With the data normalized, it’s easier to see an apparent relationship between grade and study time. It’s not an exact match, but it definitely seems like students with higher grades tend to have studied more.
So there seems to be a correlation between study time and grade; and in fact, there’s a statistical correlation measurement we can use to quantify the relationship between these columns.
df_normalized.Grade.corr(df_normalized.StudyHours)
The correlation statistic is a value between -1 and 1 that indicates the strength of a relationship. Values above 0 indicate a positive correlation (high values of one variable tend to coincide with high values of the other), while values below 0 indicate a negative correlation (high values of one variable tend to coincide with low values of the other). In this case, the correlation value is close to 1; showing a strongly positive correlation between study time and grade.
Another way to visualise the apparent correlation between two numeric columns is to use a scatter plot.
# Create a scatter plot
df_sample.plot.scatter(title='Study Time vs Grade', x='StudyHours', y='Grade')
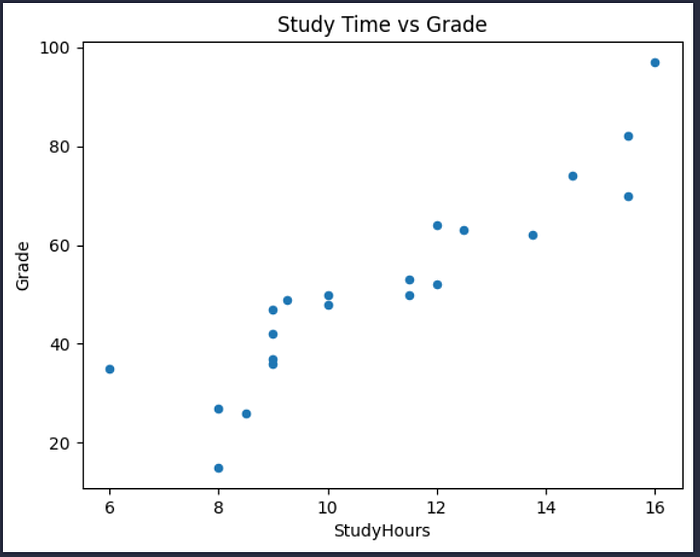
Again, it looks like there’s a discernible pattern in which the students who studied the most hours are also the students who got the highest grades.
We can see this more clearly by adding a regression line (or a line of best fit) to the plot that shows the general trend in the data. To do this, we’ll use a statistical technique called least squares regression.
Fortunately, we don’t need to code the regression calculation ourself — the SciPy package includes a stats class that provides a linregress method to do the hard work for you. This returns (among other things) the coefficients we need for the slope equation — slope (m) and intercept (b) based on a given pair of variable samples we want to compare.
from scipy import stats
#
df_regression = df_sample[['Grade', 'StudyHours']].copy()
# Get the regression slope and intercept
m, b, r, p, se = stats.linregress(df_regression['StudyHours'], df_regression['Grade'])
print('slope: {:.4f}\ny-intercept: {:.4f}'.format(m,b))
print('so...\n f(x) = {:.4f}x + {:.4f}'.format(m,b))
# Use the function (mx + b) to calculate f(x) for each x (StudyHours) value
df_regression['fx'] = (m * df_regression['StudyHours']) + b
# Calculate the error between f(x) and the actual y (Grade) value
df_regression['error'] = df_regression['fx'] - df_regression['Grade']
# Create a scatter plot of Grade vs StudyHours
df_regression.plot.scatter(x='StudyHours', y='Grade')
# Plot the regression line
plt.plot(df_regression['StudyHours'],df_regression['fx'], color='cyan')
# Display the plot
plt.show()

slope: 6.3134
y-intercept: -17.9164
so...
f(x) = 6.3134x + -17.9164
Interpretation:
- Slope (m): estimated grade increase per additional study hour
- Intercept (b): estimated grade when StudyHours = 0 (extrapolation caveat!)
Compute the Regression Line Parameters
m, b, r, p, se = stats.linregress(df_regression['StudyHours'], df_regression['Grade'])
# Show the original x,y values, the f(x) value, and the error
df_regression[['StudyHours', 'Grade', 'fx', 'error']]
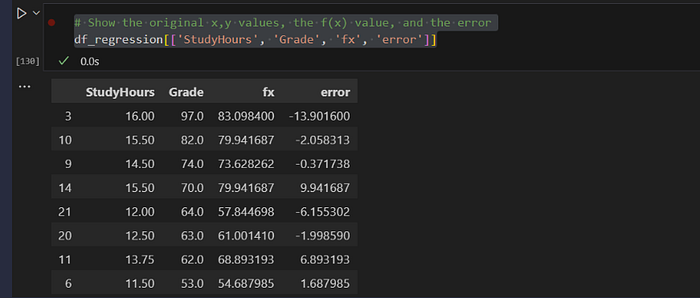
m, b, r, p, se = stats.linregress(df_regression['StudyHours'], df_regressio['Grade'])

Using the regression coefficients for prediction
Create a small predictor:
# Define a function based on our regression coefficients
def f(x):
m = 6.3134
b = -17.9164
return m*x + b
study_time = 14
# Get f(x) for study time
prediction = f(study_time)
# Grade can't be less than 0 or more than 100
expected_grade = max(0,min(100,prediction))
#Print the estimated grade
print ('Studying for {} hours per week may result in a grade of {:.0f}'.format(study_time, expected_grade))
def predict_grade(hours):
m = 6.3134
b = -17.9164
return max(0, min(100, m*hours + b))
predict_grade(14)
Studying for 14 hours per week may result in a grade of 70
Step8— Distributions & the 68–95–99.7 Rule
For roughly normal variables
- ~68.26% within 1 standard deviation of the mean
- ~95.45% within 2
- ~99.73% within 3
This helps you set expectations, detect anomalies, and communicate uncertainty.
How EDA Directly Improves Model Building
- Cleaner training data → fewer silent failures
Imputing or dropping missingness avoids brittle pipelines. - Correct feature scaling → stable training
Normalization prevents one feature from overpowering others. - Outlier handling → robust parameters
Reduces skewed loss and overfitting to rare points. - Distribution awareness → right algorithm & metrics
Skewed targets? Consider transformations or robust losses. Imbalanced classes? Choose stratified splits and the right metrics. - Relationship mapping → better feature engineering
EDA reveals interactions worth encoding and noise worth dropping. - Assumption checks → honest modeling
Linear models assume linearity and homoscedasticity; time series assume stationarity; EDA helps verify or compensate. - Leakage prevention → trustworthy validation
EDA can expose accidental target leakage or temporal leakage before you publish a suspiciously high score.
Conclusion
In this notebook, we have learned how to explore data in a Pandas DataFrame, and how to use statistical techniques to understand the relationship between variables in the sample data.
Github link:https://github.com/faizulkhan56/python-eda-discovery
Join thousands of data leaders on the AI newsletter. Join over 80,000 subscribers and keep up to date with the latest developments in AI. From research to projects and ideas. If you are building an AI startup, an AI-related product, or a service, we invite you to consider becoming a sponsor.
Published via Towards AI
Towards AI Academy
We Build Enterprise-Grade AI. We'll Teach You to Master It Too.
15 engineers. 100,000+ students. Towards AI Academy teaches what actually survives production.
Start free — no commitment:
→ 6-Day Agentic AI Engineering Email Guide — one practical lesson per day
→ Agents Architecture Cheatsheet — 3 years of architecture decisions in 6 pages
Our courses:
→ AI Engineering Certification — 90+ lessons from project selection to deployed product. The most comprehensive practical LLM course out there.
→ Agent Engineering Course — Hands on with production agent architectures, memory, routing, and eval frameworks — built from real enterprise engagements.
→ AI for Work — Understand, evaluate, and apply AI for complex work tasks.
Note: Article content contains the views of the contributing authors and not Towards AI.