MLflow Evaluation Lab — Comprehensive Guide
Author(s): Faizulkhan
Originally published on Towards AI.
Table of Contents
- Theoretical Discussion of Algorithms
- Dataset Descriptions
- Code Explanations
- Docker Deployment
- Postman Testing Guide
- Architecture Diagrams
1. Theoretical Discussion of Algorithms
1.1 Logistic Regression
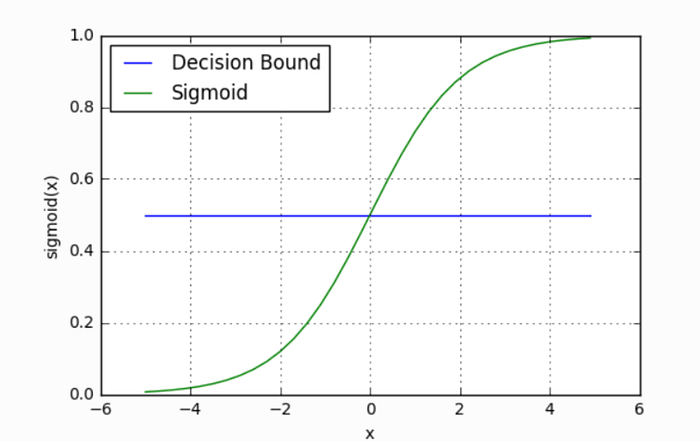
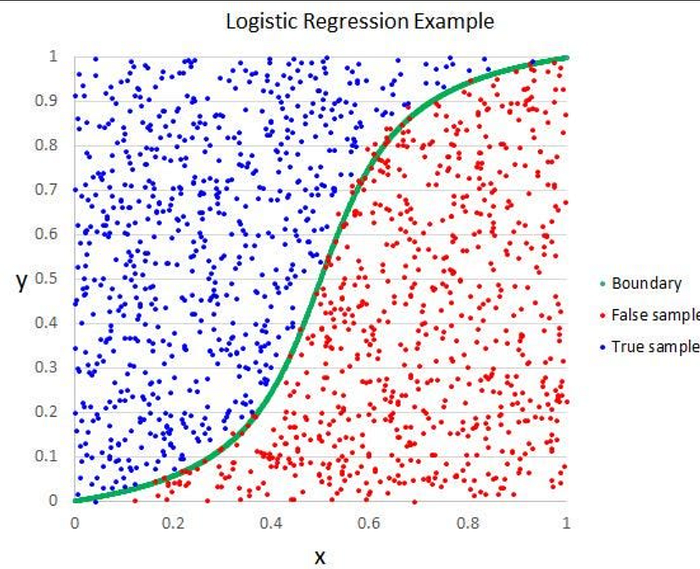
Overview: Logistic Regression is a linear classification algorithm that models the probability of a binary outcome using the logistic function (sigmoid). Despite its name, it’s used for classification, not regression.
Mathematical Foundation:
- Sigmoid Function: σ(z) = 1 / (1 + e^(-z))
- Decision Boundary: Linear boundary separating classes
- Cost Function: Binary cross-entropy loss
- Optimization: Gradient descent or variants (L-BFGS, Newton’s method)
Key Characteristics:
- Interpretability: Coefficients indicate feature importance
- Probabilistic Output: Provides probability scores, not just class labels
- Linearity: Assumes linear relationship between features and log-odds
- Regularization: L1 (Lasso) or L2 (Ridge) can prevent overfitting
Use Cases:
- Binary classification problems
- When probability estimates are needed
- Medical diagnosis (e.g., breast cancer detection)
- Credit scoring
- Spam detection
Advantages:
- Fast training and prediction
- No hyperparameter tuning required (basic version)
- Provides probability estimates
- Less prone to overfitting with regularization
Disadvantages:
- Assumes linear decision boundary
- Sensitive to outliers
- Requires feature scaling for best performance
- May underperform on non-linear problems

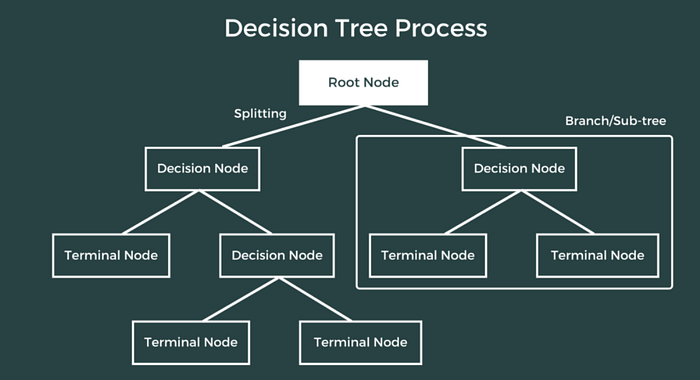
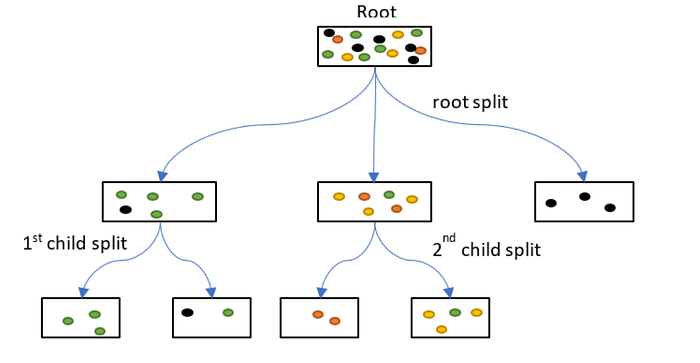
1.2 Decision Tree Regressor
Overview: Decision Tree Regressor is a non-parametric supervised learning algorithm that builds a tree-like model of decisions to predict continuous target values.
Mathematical Foundation:
- Splitting Criterion:
Mean Squared Error (MSE): Minimize variance in child nodes
Mean Absolute Error (MAE): Minimize absolute deviations
. Stopping Criteria: Maximum depth, minimum samples per leaf, minimum impurity decrease.
. Prediction: Average of target values in leaf node
Key Characteristics:
- Non-linear: Can capture complex relationships.
- Feature Selection: Automatically selects important features.
- Interpretability: Tree structure is human-readable.
- No Assumptions: No assumptions about data distribution.
Use Cases:
- Regression problems with non-linear relationships
- Feature importance analysis
- When interpretability is important
- Real estate price prediction
- Sales forecasting
Advantages:
- Easy to understand and visualize
- Handles non-linear relationships
- No feature scaling required
- Handles missing values
- Feature importance scores
Disadvantages:
- Prone to overfitting
- Sensitive to small data changes
- Can create biased trees with imbalanced classes
- May not generalize well
1.3 K-Means Clustering
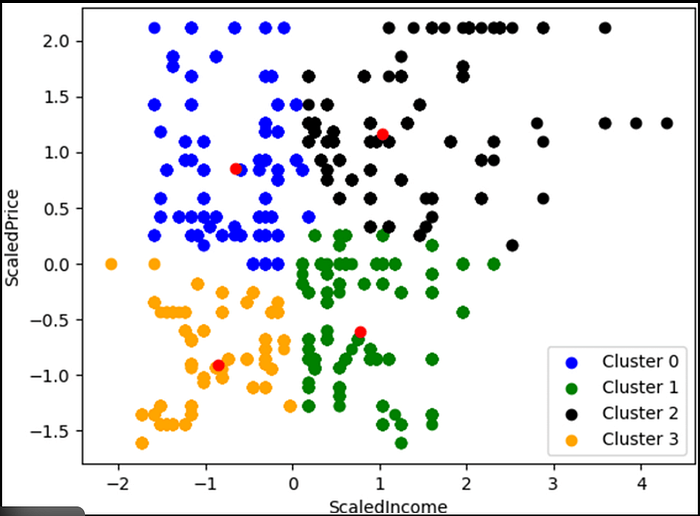
Overview: K-Means is an unsupervised learning algorithm that partitions data into k clusters by minimizing within-cluster sum of squares (WCSS).
Mathematical Foundation:
- Objective Function: Minimize Σ Σ ||x — μᵢ||²
- Where μᵢ is the centroid of cluster i
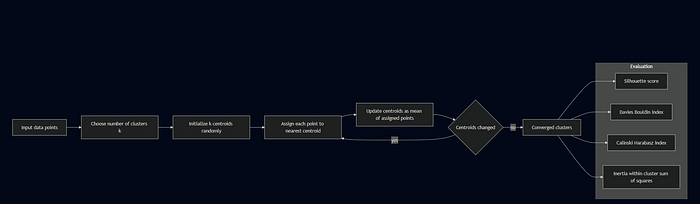
Algorithm Steps:
- Initialize k centroids randomly
- Assign each point to nearest centroid
- Update centroids as mean of assigned points
- Repeat steps 2–3 until convergence
Key Characteristics:
- Unsupervised: No labels required
- Centroid-based: Uses cluster centers
- Distance Metric: Typically Euclidean distance
- Convergence: Guaranteed to converge (may be local optimum)
Use Cases:
- Customer segmentation
- Image segmentation
- Anomaly detection
- Data exploration
- Pattern recognition
Advantages:
- Simple and fast
- Works well with spherical clusters
- Scales to large datasets
- Easy to implement
Disadvantages:
- Requires pre-specification of k
- Sensitive to initialization
- Assumes spherical clusters
- Sensitive to outliers
- May converge to local optimum
Evaluation Metrics:
- Silhouette Score: Measures how similar an object is to its own cluster vs. other clusters (-1 to 1, higher is better)
- Davies-Bouldin Index: Average similarity ratio of clusters (lower is better)
- Calinski-Harabasz Index: Ratio of between-cluster to within-cluster dispersion (higher is better)
- Inertia: Within-cluster sum of squares (lower is better)
1.4 Random Forest Classifier
Overview: Random Forest is an ensemble learning method that combines multiple decision trees to improve prediction accuracy and reduce overfitting.
Mathematical Foundation:
- Bootstrap Aggregating (Bagging): Trains multiple trees on random subsets of data
- Feature Randomness: Each split considers random subset of features
- Voting: Classification uses majority vote, regression uses average
- Out-of-Bag (OOB) Error: Validation using samples not in bootstrap
Key Characteristics:
- Ensemble Method: Combines multiple weak learners
- Randomization: Reduces overfitting through randomness
- Feature Importance: Aggregates importance across trees
- Robustness: Less sensitive to outliers than single trees
Use Cases:
- Classification with high-dimensional data
- Feature importance analysis
- When interpretability is less critical than accuracy
- Medical diagnosis
- Fraud detection
Advantages:
- Reduces overfitting compared to single trees
- Handles missing values
- Provides feature importance
- Works well with default parameters
- Handles non-linear relationships
Disadvantages:
- Less interpretable than single trees
- Can be slow for large datasets
- Memory intensive
- May overfit with noisy data
- Black box nature

Intuition
- Each tree = one expert
- Each expert gives a prediction
- Forest combines them → majority vote
- More trees → more stable answer
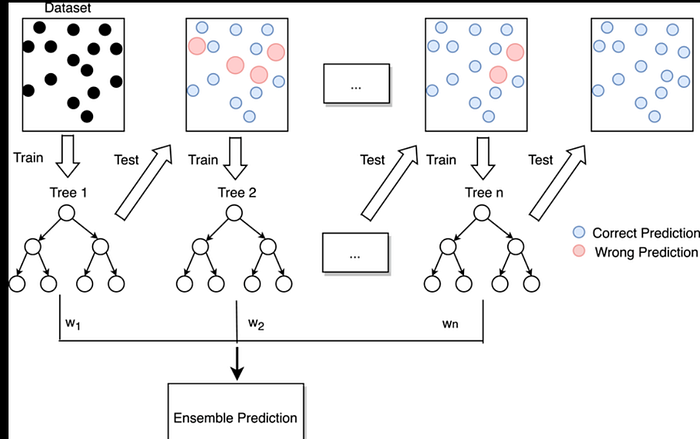
Random Forest — How It Works Internally (Tree Ensemble)

What this shows
- Input dataset is sampled into bootstrap samples
- Each tree sees a slightly different dataset
- Each node split uses random subset of features
- Final prediction = vote (classification)
1.5 Isolation Forest
Overview: Isolation Forest is an unsupervised anomaly detection algorithm that isolates anomalies instead of profiling normal data points.
Mathematical Foundation:
- Isolation Principle: Anomalies are easier to isolate (fewer splits needed)
- Isolation Tree: Binary tree where each node randomly selects feature and split value
- Path Length: Number of edges from root to leaf (anomalies have shorter paths)
- Anomaly Score: s(x, n) = 2^(-E(h(x))/c(n))
- Where E(h(x)) is average path length, c(n) is normalization constant
Key Characteristics:
- Unsupervised: No labels required
- Efficient: O(n log n) time complexity
- Scalable: Works well with high-dimensional data
- Interpretability: Can identify which features contribute to anomaly
Use Cases:
- Fraud detection
- Network intrusion detection
- Manufacturing defect detection
- Outlier detection in datasets
- Quality control
Advantages:
- Fast and efficient
- Works with high-dimensional data
- No assumptions about data distribution
- Handles multiple anomaly types
- Low memory requirement
Disadvantages:
- May struggle with high-dimensional sparse data
- Sensitive to contamination parameter
- Less interpretable than some methods
- May miss contextual anomalies
Output Interpretation:
- -1: Anomaly (outlier)
- 1: Normal (inlier)

Real intuition
- Normal points live in groups
- Anomalies are far away or isolated
- Random cuts separate anomalies faster
- Fewer splits = anomaly
How Isolation Forest Works Internally (Isolation Trees)
Isolation Forest builds many random binary trees. If a point is an anomaly, it appears near the top of most trees.
What you see
- Random feature chosen
- Random value chosen
- Split data
- Anomalies get isolated in fewer splits
Used heavily in:
- Credit card fraud
- Network intrusion detection
- Manufacturing defect analysis
Real-world mapping
- Points deviating from normal patterns
- Machines with unusual behavior
- Customers with abnormal spending
- Network traffic with strange spikes
Isolation Forest catches these because they isolate easily.
Final Output
- -1 → Anomaly
- 1 → Normal
Plot usually shows anomalies as red points far from clusters.
1.6 XGBoost Classifier
Overview: XGBoost (Extreme Gradient Boosting) is an optimized gradient boosting framework that uses ensemble of weak learners (decision trees) with gradient descent optimization.
Mathematical Foundation:
- Gradient Boosting: Sequentially adds trees that correct previous errors
- Objective Function: L(θ) = Σ l(yᵢ, ŷᵢ) + Σ Ω(fₖ)
Loss function + regularization term
- Tree Construction: Uses second-order gradient information (Hessian)
- Regularization: L1 (alpha) and L2 (lambda) regularization
Key Characteristics:
- Gradient Boosting: Sequential error correction
- Regularization: Built-in L1 and L2 regularization
- Parallel Processing: Parallel tree construction
- Handles Missing Values: Automatic handling
- Feature Importance: Multiple importance types
Use Cases:
- Classification and regression
- Kaggle competitions (often top performer)
- Large-scale machine learning
- When accuracy is critical
- Structured/tabular data
Advantages:
- High predictive accuracy
- Handles missing values automatically
- Regularization prevents overfitting
- Fast training and prediction
- Feature importance scores
Disadvantages:
- Requires hyperparameter tuning
- Less interpretable than simpler models
- Can overfit with small datasets
- Memory intensive for large datasets
- Sensitive to hyperparameters
XGBoost — Real-Life Intuition (Fixing Mistakes Step-by-Step)
XGBoost works like:
👉 “Many small trees are built one after another — each one fixes the mistakes of the previous ones.”
Intuition
- First tree makes predictions
- Next tree focuses on errors
- Next tree fixes the remaining errors
- Gradually → high accuracy
Real-Life Applications (High-Accuracy Tasks)
Use cases
- Fraud detection
- Loan approval
- Medical risk scoring
- Customer churn prediction
- Competition datasets
1.7 Linear Regressor
Overview: Linear Regression models the relationship between features and a continuous target variable using a linear equation.
Mathematical Foundation:
- Model: y = β₀ + β₁x₁ + β₂x₂ + … + βₙxₙ + ε
- Cost Function: Mean Squared Error (MSE) = (1/n) Σ(yᵢ — ŷᵢ)²
- Optimization: Ordinary Least Squares (OLS) or gradient descent
- Assumptions:
Linearity
Independence
Homoscedasticity
Normality of errors
Key Characteristics:
- Linearity: Assumes linear relationship
- Interpretability: Coefficients show feature impact
- Efficiency: Fast training and prediction
- Baseline: Often used as baseline model
Use Cases:
- Price prediction
- Sales forecasting
- Risk assessment
- When interpretability is important
- Baseline for comparison
Advantages:
- Simple and interpretable
- Fast training and prediction
- No hyperparameter tuning needed
- Provides confidence intervals
- Works well with many features
Disadvantages:
- Assumes linear relationship
- Sensitive to outliers
- Requires feature scaling
- May underfit complex relationships
- Assumes independence of features
Evaluation Metrics:
- MAE (Mean Absolute Error): Average absolute difference
- RMSE (Root Mean Squared Error): Square root of average squared difference
- R² (Coefficient of Determination): Proportion of variance explained (0 to 1, higher is better)
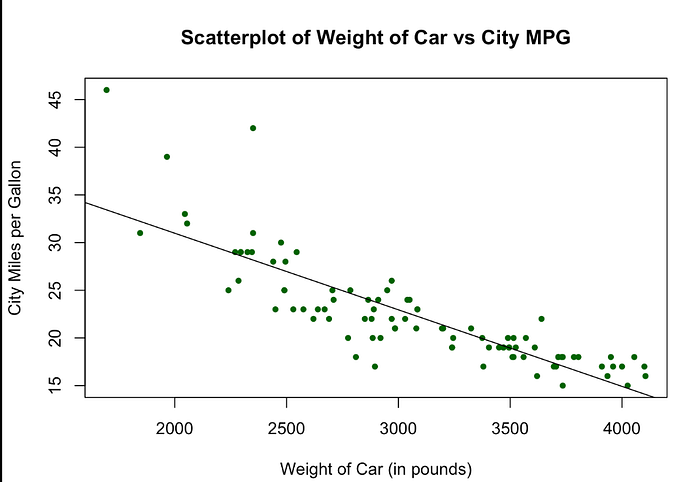
Intuition
- Data points = real observations
- Line = model prediction
- The closer points are to the line, the better the model
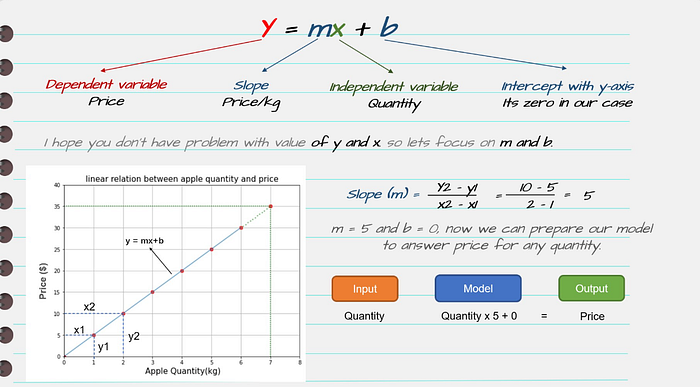
What the diagram conveys
y = b0 + b1 * x
- b0 = intercept
- b1 = slope
- x = feature
- y = prediction
Residuals — Errors Between Actual & Predicted
Residuals show how far each point is from the regression line.
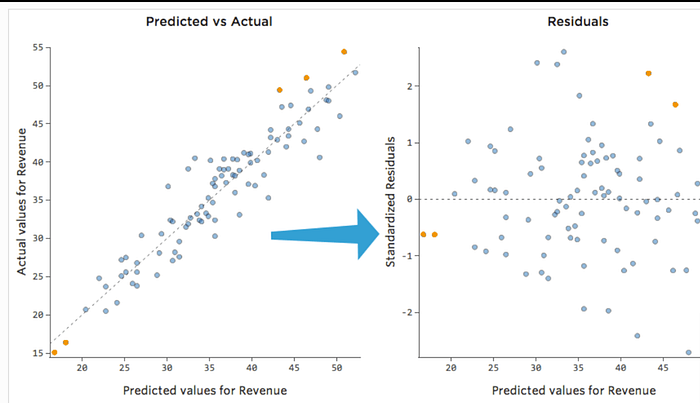
Why residuals matter
- Small, random errors → good model
- Patterns in residuals → model assumptions broken
Real-Life Use Case Diagrams
Use cases
- Predict house prices
- Forecast sales
- Analyze risk scores
Evaluation Metrics Visualized (MAE, RMSE, R²)
Understanding how Linear Regression is judged.
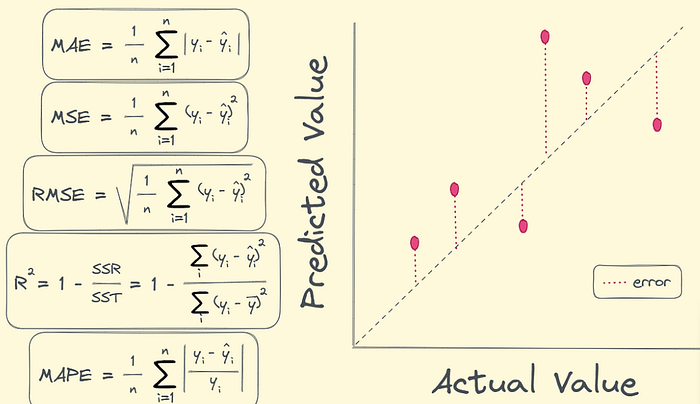
Metrics meaning
- MAE → average absolute mistake
- RMSE → penalizes large mistakes
- R² → how much of the variance your model explains
2. Dataset Descriptions
2.1 Breast Cancer Wisconsin Dataset
Source: sklearn.datasets.load_breast_cancer
Purpose: Binary classification (malignant vs. benign tumors)
Dataset Characteristics:
- Samples: 569
- Features: 30 (all numeric)
- Target: Binary (0 = malignant, 1 = benign)
- Classes: 2 (Malignant: 212, Benign: 357)
Input Features (30 features):
The dataset contains measurements of cell nuclei from breast mass images. Features are computed from digitized images and describe characteristics of cell nuclei.
Mean Features (10):
mean radius– Mean of distances from center to points on perimetermean texture– Standard deviation of gray-scale valuesmean perimeter– Mean size of the core tumormean area– Mean area of the core tumormean smoothness– Local variation in radius lengthsmean compactness– Perimeter² / area – 1.0mean concavity– Severity of concave portions of the contourmean concave points– Number of concave portions of the contourmean symmetry– Symmetry of the cellmean fractal dimension– "Coastline approximation" – 1
Standard Error Features (10): 11. radius error – Standard error of radius 12. texture error – Standard error of texture 13. perimeter error – Standard error of perimeter 14. area error – Standard error of area 15. smoothness error – Standard error of smoothness 16. compactness error – Standard error of compactness 17. concavity error – Standard error of concavity 18. concave points error – Standard error of concave points 19. symmetry error – Standard error of symmetry 20. fractal dimension error – Standard error of fractal dimension
Worst Features (10): 21. worst radius – Largest mean radius 22. worst texture – Largest standard deviation of gray-scale values 23. worst perimeter – Largest perimeter 24. worst area – Largest area 25. worst smoothness – Largest local variation 26. worst compactness – Largest compactness 27. worst concavity – Largest concavity 28. worst concave points – Largest number of concave portions 29. worst symmetry – Largest symmetry 30. worst fractal dimension – Largest fractal dimension
Target Feature:
target: Binary classification
0 = Malignant (cancerous)
1 = Benign (non-cancerous)
Used In:
- Lab 1: Logistic Regression
- Lab 5: Isolation Forest (anomaly detection)
2.2 California Housing Dataset
Source: sklearn.datasets.fetch_california_housing
Purpose: Regression (predicting median house value)
Dataset Characteristics:
- Samples: 20,640
- Features: 8 (all numeric)
- Target: Continuous (median house value in hundreds of thousands of dollars)
Input Features (8 features):
MedInc(Median Income)
- Median income in block group
- Range: ~0.5 to ~15
- Unit: Tens of thousands of dollars
HouseAge(House Age)
- Median house age in block group
- Range: 1 to 52 years
AveRooms(Average Rooms)
- Average number of rooms per household
- Range: ~0.8 to ~140
AveBedrms(Average Bedrooms)
- Average number of bedrooms per household
- Range: ~0.3 to ~50
Population(Population)
- Block group population
- Range: 3 to 35,682
AveOccup(Average Occupancy)
- Average number of household members
- Range: ~0.7 to ~1,243
Latitude(Latitude)
- Block group latitude
- Range: 32.54 to 41.95 (California)
Longitude(Longitude)
- Block group longitude
- Range: -124.35 to -114.31 (California)
Target Feature:
target: Median house value- Unit: Hundreds of thousands of dollars
- Range: ~0.15 to ~5.0
- Example: 4.21 = $421,000
Used In:
- Lab 2: Decision Tree Regressor
- Linear Regressor (regressor.py)
2.3 Iris Dataset
Source: sklearn.datasets.load_iris
Purpose: Multi-class classification (iris species identification)
Dataset Characteristics:
- Samples: 150
- Features: 4 (all numeric, in centimeters)
- Target: Multi-class (3 species)
- Classes: 3 (Setosa: 50, Versicolor: 50, Virginica: 50)
Input Features (4 features):
sepal length (cm)
- Length of sepal (outer part of flower)
- Range: 4.3 to 7.9 cm
sepal width (cm)
- Width of sepal
- Range: 2.0 to 4.4 cm
petal length (cm)
- Length of petal (inner part of flower)
- Range: 1.0 to 6.9 cm
petal width (cm)
- Width of petal
- Range: 0.1 to 2.5 cm
Target Feature:
target: Iris species0= Setosa1= Versicolor2= Virginica
Used In:
- Lab 3: K-Means Clustering (unsupervised, no labels used)
- Lab 4: Random Forest Classifier
2.4 Adult Dataset (Census Income)
Source: shap.datasets.adult() (UCI Machine Learning Repository)
Purpose: Binary classification (income prediction)
Dataset Characteristics:
- Samples: 32,561 (training) + 16,281 (test)
- Features: 14 (mix of numeric and categorical)
- Target: Binary (>50K or <=50K income)
Input Features (14 features):
Numeric Features:
age– Age of individual (17-90)fnlwgt– Final weight (sampling weight)education-num– Numeric representation of education level (1-16)capital-gain– Capital gains (0-99999)capital-loss– Capital losses (0-4356)hours-per-week– Hours worked per week (1-99)
Categorical Features: 7. workclass – Type of employment (Private, Self-emp-not-inc, etc.) 8. education – Education level (Bachelors, Some-college, etc.) 9. marital-status – Marital status (Married-civ-spouse, Divorced, etc.) 10. occupation – Occupation type (Tech-support, Craft-repair, etc.) 11. relationship – Relationship status (Husband, Wife, etc.) 12. race – Race (White, Black, Asian-Pac-Islander, etc.) 13. sex – Gender (Male, Female) 14. native-country – Country of origin (United-States, Mexico, etc.)
Target Feature:
target: Income level0= <=50K (low income)1= >50K (high income)
Used In:
- XGBoost Classifier (classification.py)
2.5 Wine Quality Red Dataset
Source: Kaggle API (uciml/red-wine-quality-cortez-et-al-2009) or UCI ML Repository
Purpose: Binary classification (wine quality prediction: good vs. bad)
Dataset Characteristics:
- Samples: 1,599
- Features: 11 (all numeric, physicochemical properties)
- Target: Binary (quality >= 7 is “good”, quality < 7 is “bad”)
- Classes: 2 (Good: ~217, Bad: ~1,382)
Input Features (11 features):
fixed acidity– Fixed acidity (tartaric acid – g/dm³)
- Range: 4.6 to 15.9
volatile acidity– Volatile acidity (acetic acid – g/dm³)
- Range: 0.12 to 1.58
citric acid– Citric acid (g/dm³)
- Range: 0.0 to 1.0
residual sugar– Residual sugar (g/dm³)
- Range: 0.9 to 15.5
chlorides– Chlorides (sodium chloride – g/dm³)
- Range: 0.012 to 0.611
free sulfur dioxide– Free sulfur dioxide (mg/dm³)
- Range: 1 to 72
total sulfur dioxide– Total sulfur dioxide (mg/dm³)
- Range: 6 to 289
density– Density (g/cm³)
- Range: 0.99007 to 1.00369
pH– pH (acidity/basicity)
- Range: 2.74 to 4.01
sulphates– Sulphates (potassium sulphate – g/dm³)
- Range: 0.33 to 2.0
alcohol– Alcohol content (% by volume)
- Range: 8.4 to 14.9
Target Feature:
quality: Wine quality score (0-10 scale)- Binary Classification:
quality >= 7= Good (1),quality < 7= Bad (0) - Original scale: 3 to 8 (no wines rated 0, 1, 2, 9, or 10 in this dataset)
Dataset Download:
- Primary Method: Kaggle API (requires credentials in
.envfile) - Fallback Method: UCI ML Repository (automatic if Kaggle unavailable)
- Helper Module:
dataset_loader.pyhandles download automatically
Used In:
- Lab 6: Wine Quality Classification with Autologging
- Lab 7: Wine Quality Classification with Manual Logging
3. Code Explanations
3.0 Dataset Loader and Kaggle Integration
Dataset Loader Module (dataset_loader.py):
The project includes a helper module for downloading datasets, particularly the Wine Quality dataset. This module supports:
- Kaggle API Integration: Downloads from Kaggle if credentials are provided
- UCI ML Repository Fallback: Automatically falls back if Kaggle is unavailable
- Local Caching: Saves dataset locally to avoid repeated downloads
Setup for Kaggle API:
- Create
.envfile in project root:
KAGGLE_USERNAME=your-username
KAGGLE_KEY=your-api-key
2. Update docker-compose.yml to read from .env:
environment:
- KAGGLE_USERNAME=${KAGGLE_USERNAME}
- KAGGLE_KEY=${KAGGLE_KEY}
3. Rebuild containers:
docker-compose build
docker-compose up -d
Usage in Lab Scripts:
from dataset_loader import load_wine_quality_dataset
data = load_wine_quality_dataset() # Automatically handles download
Benefits:
- Seamless dataset acquisition
- Works with or without Kaggle credentials
- Consistent dataset location across all scripts
- No manual download steps required
3.1 Lab 1: Logistic Regression (lab1_logistic_regression.py)
Purpose: Binary classification of breast cancer tumors (malignant vs. benign)
Code Flow:
# 1. Setup and Configuration
- Set MLflow tracking URI
- Create/select experiment: "lab1-logistic-regression"
- Load breast cancer dataset (30 features, binary target)
# 2. Data Preparation
- Split data: 80% train, 20% test
- Prepare evaluation data with target column
# 3. Model Training
- Initialize LogisticRegression (max_iter=1000, random_state=42)
- Fit model on training data
# 4. MLflow Tracking
with mlflow.start_run():
# Infer signature from test data
signature = infer_signature(X_test, y_pred)
# Log model with signature
mlflow.sklearn.log_model(model, "model", signature=signature)
# Log hyperparameters
mlflow.log_params({"max_iter": 1000, "random_state": 42, "solver": "lbfgs"})
# Evaluate model
result = mlflow_evaluate(
model=predict_fn,
data=eval_data,
targets="label",
model_type="classifier",
evaluators=["default"]
)
# Print metrics (accuracy, F1, ROC AUC, precision, recall)
Key Components:
- Model Signature: Defines input/output schema for API deployment
- MLflow Evaluate: Automatically calculates classification metrics
- Metrics Logged: Accuracy, F1 Score, ROC AUC, Precision, Recall
Evaluation Metrics Explained:
- Accuracy: (TP + TN) / (TP + TN + FP + FN) — Overall correctness
- F1 Score: 2 × (Precision × Recall) / (Precision + Recall) — Harmonic mean
- ROC AUC: Area under ROC curve — Model’s ability to distinguish classes
- Precision: TP / (TP + FP) — Of predicted positives, how many are correct
- Recall: TP / (TP + FN) — Of actual positives, how many were found
3.2 Lab 2: Decision Tree Regressor (lab2_decision_tree_regressor.py)
Purpose: Predict median house value using decision tree regression
Code Flow:
# 1. Setup
- Experiment: "lab2-decision-tree-regressor"
- Load California Housing dataset (8 features, continuous target)
# 2. Data Preparation
- Split: 80% train, 20% test
- Prepare evaluation data
# 3. Model Training
- DecisionTreeRegressor(max_depth=10, random_state=42)
- Fit on training data
# 4. MLflow Tracking
with mlflow.start_run():
# Infer signature
signature = infer_signature(X_test, y_pred)
# Log model
mlflow.sklearn.log_model(model, "model", signature=signature)
# Log parameters
mlflow.log_params({
"max_depth": 10,
"random_state": 42,
"criterion": "squared_error"
})
# Evaluate model
result = mlflow_evaluate(
model=predict_fn,
data=eval_data,
targets="target",
model_type="regressor",
evaluators=["default"]
)
# Print metrics (MAE, RMSE, R²)
Key Components:
- Max Depth: Limits tree depth to prevent overfitting
- Regression Metrics: MAE, RMSE, R² score
- Feature Importance: Decision trees provide feature importance scores
Evaluation Metrics Explained:
- MAE (Mean Absolute Error): Average absolute difference between predicted and actual
- RMSE (Root Mean Squared Error): Square root of average squared differences (penalizes large errors)
- R² (R-squared): Proportion of variance explained (1.0 = perfect, 0.0 = no better than mean)
3.3 Lab 3: K-Means Clustering (lab3_kmeans_clustering.py)
Purpose: Unsupervised clustering of iris flowers into 3 groups
Code Flow:
# 1. Setup
- Experiment: "lab3-kmeans-clustering"
- Load Iris dataset (features only, no labels for clustering)
# 2. Data Preparation
- Use all data (no train/test split for unsupervised learning)
- Sample for signature inference
# 3. Model Training
- KMeans(n_clusters=3, random_state=42, n_init=10)
- Fit on all data
# 4. MLflow Tracking
with mlflow.start_run():
# Predict clusters
y_pred = model.predict(X)
# Infer signature
signature = infer_signature(X_sample, model.predict(X_sample))
# Log model
mlflow.sklearn.log_model(model, "model", signature=signature)
# Log parameters
mlflow.log_params({
"n_clusters": 3,
"random_state": 42,
"n_init": 10
})
# Calculate clustering metrics manually
silhouette = silhouette_score(X, y_pred)
davies_bouldin = davies_bouldin_score(X, y_pred)
calinski_harabasz = calinski_harabasz_score(X, y_pred)
inertia = model.inertia_
# Log metrics
mlflow.log_metrics({
"silhouette_score": silhouette,
"davies_bouldin_score": davies_bouldin,
"calinski_harabasz_score": calinski_harabasz,
"inertia": inertia
})
Key Components:
- Unsupervised Learning: No labels used during training
- Manual Metrics: MLflow doesn’t have built-in clustering evaluators
- Cluster Validation: Multiple metrics assess cluster quality
Clustering Metrics Explained:
- Silhouette Score: Measures how similar objects are to their cluster vs. others (-1 to 1)
- Davies-Bouldin Index: Average similarity ratio (lower is better)
- Calinski-Harabasz Index: Ratio of between-cluster to within-cluster dispersion (higher is better)
- Inertia: Within-cluster sum of squares (lower is better)
3.4 Lab 4: Random Forest Classifier (lab4_random_forest_classifier.py)
Purpose: Multi-class classification of iris species using ensemble of trees
Code Flow:
# 1. Setup
- Experiment: "lab4-random-forest-classifier"
- Load Iris dataset (4 features, 3 classes)
# 2. Data Preparation
- Split: 80% train, 20% test
- Prepare evaluation data
# 3. Model Training
- RandomForestClassifier(n_estimators=100, max_depth=5, random_state=42)
- Fit on training data
# 4. MLflow Tracking
with mlflow.start_run():
# Infer signature
signature = infer_signature(X_test, y_pred)
# Log model
mlflow.sklearn.log_model(model, "model", signature=signature)
# Log parameters
mlflow.log_params({
"n_estimators": 100,
"max_depth": 5,
"random_state": 42
})
# Evaluate model
result = mlflow_evaluate(
model=predict_fn,
data=eval_data,
targets="label",
model_type="classifier",
evaluators=["default"]
)
# Print metrics (accuracy, F1, precision, recall per class)
Key Components:
- Ensemble Method: Combines 100 decision trees
- Multi-class Classification: Handles 3 classes (setosa, versicolor, virginica)
- Feature Importance: Aggregates importance across all trees
3.5 Lab 5: Isolation Forest (lab5_isolation_forest.py)
Purpose: Anomaly detection on breast cancer dataset
Code Flow:
# 1. Setup
- Experiment: "lab5-isolation-forest"
- Load breast cancer dataset (30 features)
# 2. Data Preparation
- Use all data (unsupervised)
- Sample for signature inference
# 3. Model Training
- IsolationForest(contamination=0.1, random_state=42)
- Fit on all data
# 4. MLflow Tracking
with mlflow.start_run():
# Predict anomalies
y_pred = model.predict(X)
# Infer signature
signature = infer_signature(X_sample, model.predict(X_sample))
# Log model
mlflow.sklearn.log_model(model, "model", signature=signature)
# Log parameters
mlflow.log_params({
"contamination": 0.1,
"random_state": 42,
"n_estimators": 100
})
# Calculate metrics manually
n_anomalies = (y_pred == -1).sum()
n_normal = (y_pred == 1).sum()
anomaly_ratio = n_anomalies / len(y_pred)
# Log metrics
mlflow.log_metrics({
"n_anomalies": n_anomalies,
"n_normal": n_normal,
"anomaly_ratio": anomaly_ratio
})
Key Components:
- Contamination Parameter: Expected proportion of anomalies (0.1 = 10%)
- Anomaly Detection: Returns -1 (anomaly) or 1 (normal)
- Unsupervised: No labels required
3.6 XGBoost Classifier (classification.py)
Purpose: Binary classification on Adult dataset with SHAP explainability
Code Flow:
# 1. Setup
- Experiment: "evaluation-classification"
- Load Adult dataset from SHAP
# 2. Data Preparation
- Split: 80% train, 20% test
- XGBoost handles categorical features automatically
# 3. Model Training
- XGBClassifier() with default parameters
- Fit on training data
# 4. MLflow Tracking
with mlflow.start_run():
# Infer signature
signature = infer_signature(X_test, y_pred)
# Log model
mlflow.sklearn.log_model(model, "model", signature=signature)
# Evaluate model
result = mlflow_evaluate(...)
# SHAP Explainability
explainer = shap.TreeExplainer(model)
shap_values = explainer.shap_values(X_test[:100])
# Create SHAP plots
shap.summary_plot(shap_values, X_test[:100], show=False)
plt.savefig("shap_summary.png")
# Log SHAP plot as artifact
mlflow.log_artifact("shap_summary.png")
Key Components:
- SHAP (SHapley Additive exPlanations): Explains model predictions
- Feature Importance: Shows which features drive predictions
- Artifact Logging: Saves SHAP plots to MLflow
3.7 Linear Regressor (regressor.py)
Purpose: Predict median house value using linear regression
Code Flow:
# 1. Setup
- Experiment: "evaluation-regression"
- Load California Housing dataset
# 2. Data Preparation
- Split: 80% train, 20% test
- Prepare evaluation data
# 3. Model Training
- LinearRegression()
- Fit on training data
# 4. MLflow Tracking
with mlflow.start_run():
# Infer signature
signature = infer_signature(X_test, y_pred)
# Log model
mlflow.sklearn.log_model(model, "model", signature=signature)
# Evaluate model
result = mlflow_evaluate(
model=predict_fn,
data=eval_data,
targets="target",
model_type="regressor",
evaluators=["default"]
)
# Print metrics (MAE, RMSE, R²)
3.8 Dataset Loader (dataset_loader.py)
Purpose: Helper module for downloading Wine Quality dataset from Kaggle or UCI ML Repository
Key Features:
- Automatic dataset download with fallback mechanism
- Supports Kaggle API (if credentials provided) or UCI ML Repository
- Handles file extraction and path management
- Caches dataset locally to avoid repeated downloads
Code Flow:
def load_wine_quality_dataset(data_file="data/winequality-red.csv"):
# 1. Check if local file exists
if os.path.exists(data_file):
return pd.read_csv(data_file, sep=";")
# 2. Try Kaggle API (if credentials available)
if KAGGLE_USERNAME and KAGGLE_KEY:
try:
api = KaggleApi()
api.authenticate()
api.dataset_download_files(
"uciml/red-wine-quality-cortez-et-al-2009",
path="data",
unzip=True
)
# Find and return CSV file
except Exception:
# Fallback to UCI
pass
# 3. Fallback to UCI ML Repository
data_url = "https://archive.ics.uci.edu/ml/..."
data = pd.read_csv(data_url, sep=";")
data.to_csv(data_file, index=False)
return data
Usage:
from dataset_loader import load_wine_quality_dataset
data = load_wine_quality_dataset()
Configuration:
- Kaggle credentials: Set
KAGGLE_USERNAMEandKAGGLE_KEYin.envfile - Dataset location:
data/winequality-red.csv(created automatically)
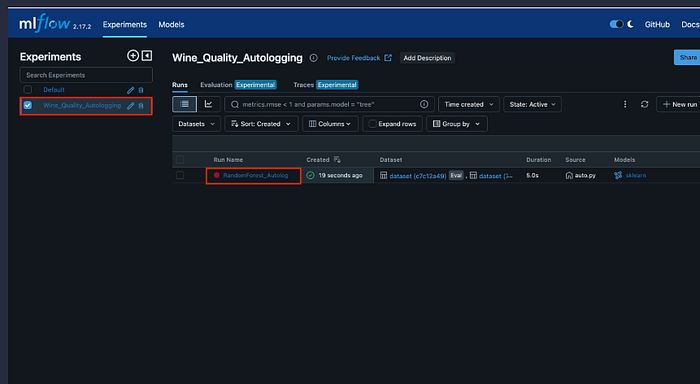
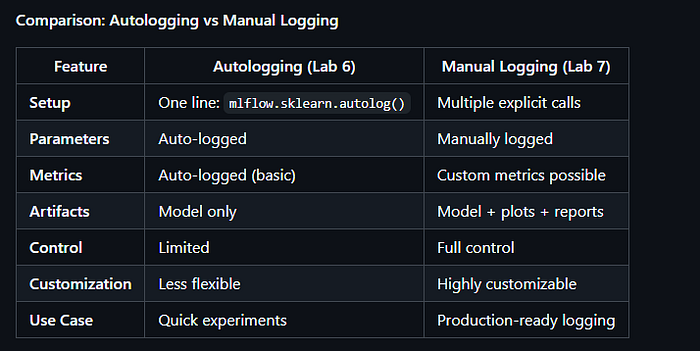
3.9 Lab 6: Wine Quality Classification with Autologging (lab6_wine_quality_autologging.py)
Purpose: Binary classification of wine quality using Random Forest with MLflow autologging.
Key Feature: Demonstrates MLflow’s autologging capability, which automatically logs parameters, metrics, and models without manual intervention.
Code Flow:
# 1. Setup and Configuration
- Set MLflow tracking URI
- Create/select experiment: "lab6-wine-quality-autologging"
- Load Wine Quality dataset using dataset_loader
# 2. Data Preparation
- Load dataset (downloads from Kaggle or UCI if needed)
- Binary classification: quality >= 7 is "good" (1), else "bad" (0)
- Split: 80% train, 20% test
# 3. Enable Autologging
mlflow.sklearn.autolog() # Enable automatic logging
# 4. Model Training (Autologging Active)
with mlflow.start_run(run_name="random_forest_autolog"):
# Train Random Forest
model = RandomForestClassifier(
n_estimators=100,
max_depth=5,
random_state=42
)
model.fit(X_train, y_train)
# Autologging automatically logs:
# - Model parameters (n_estimators, max_depth, etc.)
# - Training metrics (if available)
# - Model artifacts
# - Model signature
# Generate predictions
y_pred = model.predict(X_test)
accuracy = accuracy_score(y_test, y_pred)
# Infer signature explicitly
signature = infer_signature(X_test, y_pred)
# Log model with signature
mlflow.sklearn.log_model(model, "model", signature=signature)
# MLflow evaluation for additional metrics
result = mlflow_evaluate(...)
# 5. Disable Autologging
mlflow.sklearn.autolog(disable=True)
Key Components:
- Autologging: Automatically captures model parameters, metrics, and artifacts
- Dataset Integration: Uses
dataset_loader.pyfor automatic dataset download - Kaggle Support: Downloads from Kaggle if credentials are available in
.env - Fallback Mechanism: Automatically falls back to UCI ML Repository if Kaggle unavailable
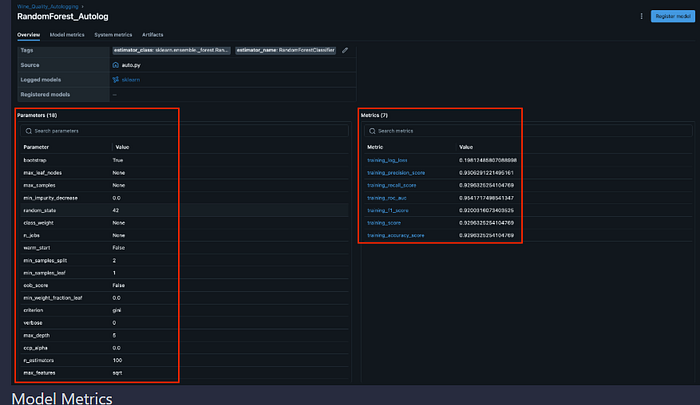
What Autologging Logs Automatically:
- Model hyperparameters (n_estimators, max_depth, random_state, etc.)
- Model signature (input/output schema)
- Model artifacts (saved model files)
- Training time and metadata
- Additional sklearn-specific metrics (if available)
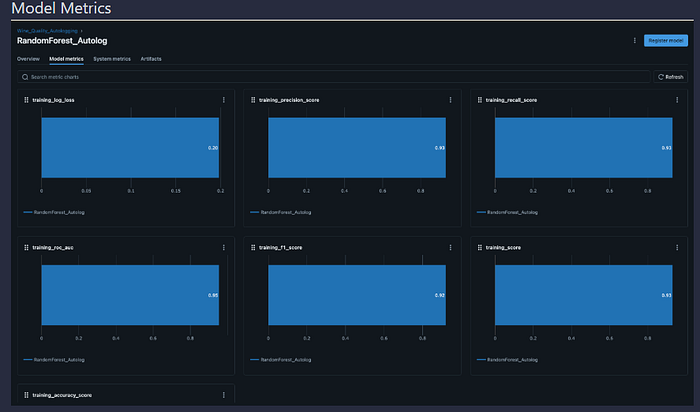
Evaluation Metrics:
- Accuracy, F1 Score, ROC AUC, Precision, Recall (via MLflow evaluators)
Running the Lab:
# With Docker
docker-compose exec mlflow-app python lab6_wine_quality_autologging.py
# Prerequisites: .env file with Kaggle credentials (optional)
# KAGGLE_USERNAME=your-username
# KAGGLE_KEY=your-api-key
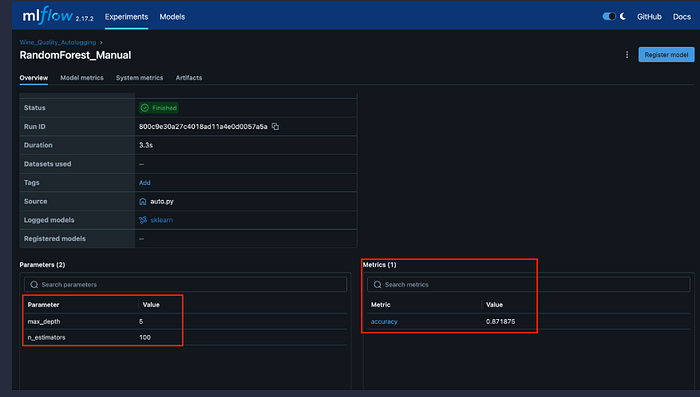
3.10 Lab 7: Wine Quality Classification with Manual Logging (lab7_wine_quality_manual.py)
Purpose: Binary classification of wine quality using Random Forest with manual MLflow logging
Key Feature: Demonstrates explicit manual logging of parameters, metrics, artifacts, and models, providing full control over what gets logged.
Code Flow:
# 1. Setup and Configuration
- Set MLflow tracking URI
- Create/select experiment: "lab7-wine-quality-manual"
- Load Wine Quality dataset using dataset_loader
# 2. Data Preparation
- Load dataset (downloads from Kaggle or UCI if needed)
- Binary classification: quality >= 7 is "good" (1), else "bad" (0)
- Split: 80% train, 20% test
# 3. Model Training
model = RandomForestClassifier(
n_estimators=100,
max_depth=5,
random_state=42,
criterion="gini"
)
model.fit(X_train, y_train)
# 4. Manual Logging
with mlflow.start_run(run_name="random_forest_manual"):
# Manual logging: Parameters
mlflow.log_params({
"n_estimators": 100,
"max_depth": 5,
"random_state": 42,
"criterion": "gini"
})
# Calculate metrics
y_pred = model.predict(X_test)
accuracy = accuracy_score(y_test, y_pred)
cm = confusion_matrix(y_test, y_pred)
precision = tp / (tp + fp)
recall = tp / (tp + fn)
f1 = 2 * (precision * recall) / (precision + recall)
# Manual logging: Metrics
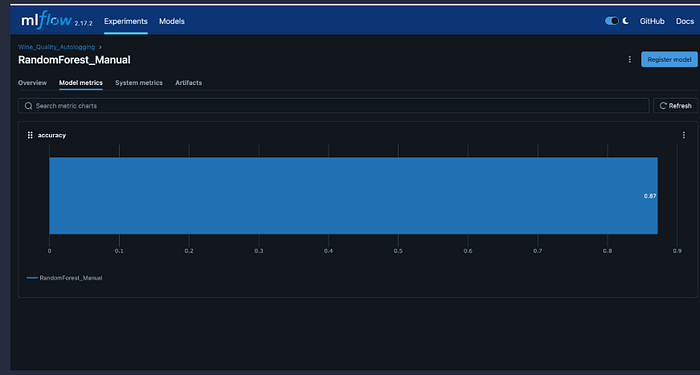
mlflow.log_metric("accuracy", accuracy)
mlflow.log_metric("precision", precision)
mlflow.log_metric("recall", recall)
mlflow.log_metric("f1_score", f1)
mlflow.log_metric("true_positives", float(tp))
mlflow.log_metric("true_negatives", float(tn))
mlflow.log_metric("false_positives", float(fp))
mlflow.log_metric("false_negatives", float(fn))
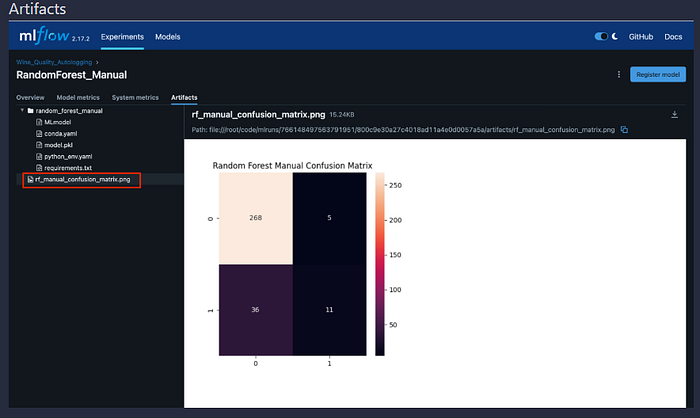
# Manual logging: Confusion Matrix as artifact
plt.figure(figsize=(8, 6))
sns.heatmap(cm, annot=True, fmt="d", cmap="Blues")
plt.savefig("rf_manual_confusion_matrix.png")
mlflow.log_artifact("rf_manual_confusion_matrix.png")
# Manual logging: Classification Report as artifact
report = classification_report(y_test, y_pred, output_dict=True)
report_df = pd.DataFrame(report).transpose()
report_df.to_csv("rf_manual_classification_report.csv")
mlflow.log_artifact("rf_manual_classification_report.csv")
# Manual logging: Feature Importance Plot as artifact
feature_importance = pd.DataFrame({
"feature": X.columns,
"importance": model.feature_importances_
}).sort_values("importance", ascending=False)
# Create and save plot
mlflow.log_artifact("rf_manual_feature_importance.png")
# Manual logging: Model
signature = infer_signature(X_test, y_pred)
mlflow.sklearn.log_model(model, "model", signature=signature)
# Manual logging: Dataset info
mlflow.log_param("dataset_size", len(data))
mlflow.log_param("train_size", len(X_train))
mlflow.log_param("test_size", len(X_test))
mlflow.log_param("n_features", len(X.columns))
# MLflow evaluation for additional metrics
result = mlflow_evaluate(...)
Key Components:
- Manual Parameter Logging: Explicitly logs all hyperparameters
- Manual Metric Logging: Calculates and logs custom metrics
- Artifact Logging: Saves plots and reports as artifacts
Confusion matrix heatmap
Classification report (CSV)
Feature importance plot
- Dataset Integration: Uses
dataset_loader.pyfor automatic dataset download - Kaggle Support: Downloads from Kaggle if credentials are available in
.env
What Gets Logged Manually:
- Model hyperparameters (via
mlflow.log_params()) - Evaluation metrics (via
mlflow.log_metric()) - Visualization artifacts (confusion matrix, feature importance plots)
- Data artifacts (classification reports)
- Model file (via
mlflow.sklearn.log_model()) - Dataset metadata (size, splits, feature count)
Evaluation Metrics:
- Accuracy, Precision, Recall, F1 Score (manually calculated)
- Additional metrics via MLflow evaluators (ROC AUC, etc.)
Running the Lab:
# With Docker
docker-compose exec mlflow-app python lab7_wine_quality_manual.py
# Prerequisites: .env file with Kaggle credentials (optional)
# KAGGLE_USERNAME=your-username
# KAGGLE_KEY=your-api-key



3.11 FastAPI Prediction Service (api_server.py)
Purpose: RESTful API for real-time model predictions
Architecture:
# 1. Application Setup
- FastAPI app with CORS middleware
- MLflow tracking URI configuration
- Global model cache dictionary
# 2. Model Loading
- Background loading on startup (non-blocking)
- Lazy loading on first request if not loaded
- Error handling for missing models
# 3. Feature Mapping
- map_breast_cancer_features(): Maps API names to sklearn names
- Handles underscore vs. space differences
# 4. Prediction Endpoints
- /predict/lab1-logistic-regression
- /predict/lab2-decision-tree-regressor
- /predict/lab3-kmeans-clustering
- /predict/lab4-random-forest
- /predict/lab5-isolation-forest
- /predict/xgboost-classifier
- /predict/linear-regressor
# 5. Health Check
- /health: Returns status and loaded models
Key Functions:
load_model_from_latest_run():
- Retrieves latest run from MLflow experiment
- Loads model using MLflow’s sklearn loader
- Handles experiment/run not found errors
ensure_model_loaded():
- Checks if model is in cache
- Loads model if missing (lazy loading)
- Prevents duplicate loading attempts
map_breast_cancer_features():
- Converts API feature names (underscores) to sklearn names (spaces)
- Required for Breast Cancer and Isolation Forest models
Request/Response Flow:
Client Request
↓
FastAPI Endpoint
↓
Pydantic Validation (feature names, types)
↓
Feature Mapping (if needed)
↓
Model Prediction
↓
Response Formatting
↓
JSON Response
Error Handling:
- 503 Service Unavailable: Model not loaded or currently loading
- 500 Internal Server Error: Prediction error (feature mismatch, etc.)
- Clear error messages with troubleshooting hints
4. Docker Deployment
4.1 Docker Compose Architecture
Services:
- PostgreSQL (
postgres)
- Image:
postgres:15-alpine - Port: 5432
- Database:
mlflow - User:
mlflow/ Password:mlflow - Volume:
postgres-data(persistent)
- MLflow Server (
mlflow-server)
- Built from:
Dockerfile.mlflow-server - Port: 5000
- Depends on: PostgreSQL (health check)
- Volume:
mlflow-artifacts(shared with app and API)
- MLflow App for training and evaluation model trainning service(
mlflow-app)
- Built from:
Dockerfile - Purpose: Run evaluation scripts
- Depends on: MLflow Server (health check)
- Volumes:
mlflow-artifacts+ project directory (bind mount)
- MLflow API for prediction servic (
mlflow-api)
- Built from:
Dockerfile.api - Port: 8000
- Depends on: MLflow Server (health check)
- Volume:
mlflow-artifacts(for model loading)
Volumes:
postgres-data: PostgreSQL database filesmlflow-artifacts: MLflow model artifacts and runs
4.2 Dockerfile.mlflow-server
Purpose: MLflow tracking server with PostgreSQL support
FROM python:3.11-slim
# Install system dependencies
RUN apt-get update && \
apt-get install -y --no-install-recommends \
postgresql-client curl && \
rm -rf /var/lib/apt/lists/*
# Install Python dependencies
RUN pip install --upgrade pip && \
pip install --no-cache-dir \
mlflow>=2.0.0 \
psycopg2-binary>=2.9.0
# Copy entrypoint script
COPY entrypoint-mlflow.sh /app/entrypoint-mlflow.sh
RUN chmod +x /app/entrypoint-mlflow.sh
WORKDIR /app
# Use entrypoint script
ENTRYPOINT ["/app/entrypoint-mlflow.sh"]
Key Components:
postgresql-client: Providespg_isreadyfor health checkspsycopg2-binary: PostgreSQL adapter for Pythonentrypoint-mlflow.sh: Custom startup script
4.3 entrypoint-mlflow.sh
Purpose: Wait for PostgreSQL and start MLflow server
#!/bin/bash
set -e
# Wait for PostgreSQL
until pg_isready -h "$POSTGRES_HOST" -p "$POSTGRES_PORT" -U "$POSTGRES_USER"; do
echo "Waiting for PostgreSQL..."
sleep 2
done
# Construct connection URI
export MLFLOW_BACKEND_STORE_URI="postgresql://${POSTGRES_USER}:${POSTGRES_PASSWORD}@${POSTGRES_HOST}:${POSTGRES_PORT}/${POSTGRES_DB}"
# Start MLflow server
exec mlflow server \
--host 0.0.0.0 \
--port 5000 \
--backend-store-uri "$MLFLOW_BACKEND_STORE_URI" \
--default-artifact-root /app/mlruns
Key Features:
- Waits for PostgreSQL to be ready
- Constructs PostgreSQL connection URI from environment variables
- Starts MLflow server with proper configuration
4.4 Dockerfile
Purpose: Python environment for running evaluation scripts
FROM python:3.11-slim
# Install system dependencies
RUN apt-get update && \
apt-get install -y --no-install-recommends gcc g++ && \
rm -rf /var/lib/apt/lists/*
# Copy requirements and install
COPY requirements.txt .
COPY pip.conf /etc/pip.conf
RUN pip install --upgrade pip && \
pip install --no-cache-dir --default-timeout=100 --retries 5 -r requirements.txt
WORKDIR /app
# Keep container running
CMD ["tail", "-f", "/dev/null"]
Key Components:
gcc,g++: Required for compiling some Python packagespip.conf: Configuration for timeouts and retries- Keeps container running for manual script execution
4.5 Dockerfile.api
Purpose: FastAPI server with model dependencies
FROM python:3.11-slim
# Install system dependencies
RUN apt-get update && \
apt-get install -y --no-install-recommends gcc g++ curl && \
rm -rf /var/lib/apt/lists/*
# Install Python dependencies in batches
RUN pip install --upgrade pip && \
pip install --no-cache-dir --default-timeout=100 --retries 5 \
mlflow>=2.0.0 scikit-learn>=1.0.0 && \
pip install --no-cache-dir --default-timeout=100 --retries 5 \
xgboost>=2.0.0 shap>=0.40.0 && \
pip install --no-cache-dir --default-timeout=100 --retries 5 \
pandas>=1.5.0 numpy>=1.24.0 psycopg2-binary>=2.9.0 && \
pip install --no-cache-dir --default-timeout=100 --retries 5 \
fastapi>=0.104.0 uvicorn[standard]>=0.24.0 pydantic>=2.0.0 requests>=2.31.0
# Copy API server
COPY api_server.py .
EXPOSE 8000
# Set environment variables
ENV MLFLOW_TRACKING_URI=http://mlflow-server:5000
ENV MLFLOW_REGISTRY_URI=http://mlflow-server:5000
ENV GIT_PYTHON_REFRESH=quiet
# Run FastAPI server
CMD ["uvicorn", "api_server:app", "--host", "0.0.0.0", "--port", "8000"]
Key Components:
- Batch installation: Prevents timeout on large packages
- All model dependencies: MLflow, scikit-learn, XGBoost, SHAP
- FastAPI stack: FastAPI, Uvicorn, Pydantic
- Health check: Uses curl for health endpoint
4.6 Deployment Steps
- Build Images:
docker-compose build
2. Start Services:
docker-compose up -d
3. Verify Services:
docker-compose ps
4. Check Logs:
# All services
docker-compose logs -f
# Specific service
docker-compose logs -f mlflow-api
5. Run Experiments:
# Individual
docker-compose exec mlflow-app python lab1_logistic_regression.py
# All experiments
docker-compose exec mlflow-app bash -c "./run_all_experiments.sh"
6. Access Services:
- MLflow UI: http://localhost:5000
- FastAPI Docs: http://localhost:8000/docs
- FastAPI Health: http://localhost:8000/health

7. Stop Services:
docker-compose down
8. Clean Up (⚠️ Deletes Data):
docker-compose down -v
4.7 Health Checks
PostgreSQL:
healthcheck:
test: ["CMD-SHELL", "pg_isready -U mlflow"]
interval: 5s
timeout: 5s
retries: 5
MLflow Server:
healthcheck:
test: ["CMD", "curl", "-f", "http://127.0.0.1:5000/health"]
interval: 10s
timeout: 5s
retries: 10
start_period: 30s
FastAPI:
healthcheck:
test: ["CMD", "curl", "-f", "http://localhost:8000/health"]
interval: 10s
timeout: 5s
retries: 10
start_period: 600s # 10 minutes for model loading
4.8 Network Configuration
Docker Network:
- All services on same network (default)
- Service names as hostnames:
postgres→ PostgreSQLmlflow-server→ MLflow Tracking Servermlflow-app→ Python Application Containermlflow-api→ FastAPI Prediction Server
Service Communication:
- Services communicate via Docker’s internal network
- No need to expose internal ports (only external ports are exposed)
- Environment variables configure service discovery
Port Mapping:
- PostgreSQL:
5432:5432(host:container) - MLflow Server:
5000:5000 - FastAPI:
8000:8000
4.9 Volume Management
Persistent Volumes:
- postgres-data:
- Stores PostgreSQL database files
- Persists experiments, runs, metrics, parameters
- Survives container restarts and removals
- Location: Docker volume directory
- mlflow-artifacts:
- Stores MLflow model artifacts (saved models, plots, etc.)
- Shared between
mlflow-server,mlflow-app, andmlflow-api - Ensures models are accessible across services
- Location:
/app/mlrunsin containers
Volume Commands:
# List volumes
docker volume ls
# Inspect volume
docker volume inspect mlflow_evaluation_lab_postgres-data
# Remove volumes (⚠️ Deletes all data)
docker-compose down -v
4.10 Environment Variables
PostgreSQL Service:
POSTGRES_USER: Database user (default:mlflow)POSTGRES_PASSWORD: Database password (default:mlflow)POSTGRES_DB: Database name (default:mlflow)
MLflow Server:
POSTGRES_HOST: PostgreSQL hostname (default:postgres)POSTGRES_PORT: PostgreSQL port (default:5432)POSTGRES_USER: Database userPOSTGRES_PASSWORD: Database passwordPOSTGRES_DB: Database name
MLflow App:
MLFLOW_TRACKING_URI: MLflow server URL (default:http://mlflow-server:5000)MLFLOW_REGISTRY_URI: MLflow registry URL (default:http://mlflow-server:5000)
MLflow API:
MLFLOW_TRACKING_URI: MLflow server URLMLFLOW_REGISTRY_URI: MLflow registry URLGIT_PYTHON_REFRESH: Suppress Git warnings (set toquiet)
4.11 Troubleshooting Docker Deployment
Issue: Services won’t start
# Check service status
docker-compose ps
# Check logs
docker-compose logs
# Restart services
docker-compose restart
Issue: Models not loading in API
- Ensure models are trained first: Run all lab scripts
- Check MLflow UI: Verify experiments and runs exist
- Check API logs:
docker-compose logs mlflow-api - Verify model loading:
curl http://localhost:8000/health
Issue: PostgreSQL connection errors
- Wait for PostgreSQL to be healthy: Check
docker-compose ps - Verify environment variables in
docker-compose.yml - Check PostgreSQL logs:
docker-compose logs postgres
Issue: Volume permission errors
- On Linux: Ensure Docker has proper permissions
- On Windows: Use Docker Desktop with WSL2 backend
- Check volume mounts:
docker volume inspect <volume-name>
5. Postman Testing Guide
5.1 Setup
Prerequisites:
- Docker services running (
docker-compose up -d) - All models trained (run all lab scripts)
- Postman installed (or use cURL)
Base URL:
http://localhost:8000
API Documentation:
- Swagger UI:
http://localhost:8000/docs - ReDoc:
http://localhost:8000/redoc
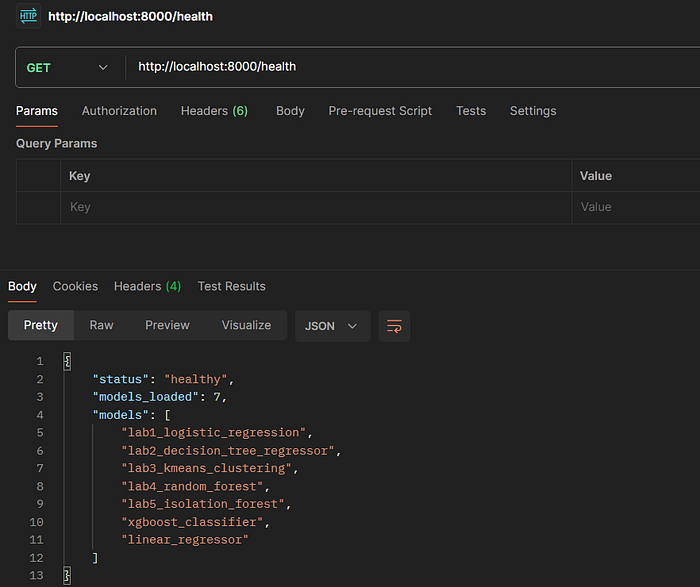
5.2 Health Check Endpoint
Request:
GET http://localhost:8000/health
Expected Response:
{
"status": "healthy",
"models_loaded": 7,
"models": [
"lab1_logistic_regression",
"lab2_decision_tree_regressor",
"lab3_kmeans_clustering",
"lab4_random_forest",
"lab5_isolation_forest",
"xgboost_classifier",
"linear_regressor"
]
}
Postman Steps:
- Create new request
- Method:
GET - URL:
http://localhost:8000/health - Click “Send”
- Verify
models_loadedequals 7
5.3 Lab 1: Logistic Regression (Breast Cancer)
Endpoint: POST /predict/lab1-logistic-regression
Request Body:
{
"mean_radius": 17.99,
"mean_texture": 10.38,
"mean_perimeter": 122.8,
"mean_area": 1001.0,
"mean_smoothness": 0.1184,
"mean_compactness": 0.2776,
"mean_concavity": 0.3001,
"mean_concave_points": 0.1471,
"mean_symmetry": 0.2419,
"mean_fractal_dimension": 0.07871,
"radius_error": 1.095,
"texture_error": 0.9053,
"perimeter_error": 8.589,
"area_error": 153.4,
"smoothness_error": 0.006399,
"compactness_error": 0.04904,
"concavity_error": 0.05373,
"concave_points_error": 0.01587,
"symmetry_error": 0.03003,
"fractal_dimension_error": 0.006193,
"worst_radius": 25.38,
"worst_texture": 17.33,
"worst_perimeter": 184.6,
"worst_area": 2019.0,
"worst_smoothness": 0.1622,
"worst_compactness": 0.6656,
"worst_concavity": 0.7119,
"worst_concave_points": 0.2654,
"worst_symmetry": 0.4601,
"worst_fractal_dimension": 0.1189
}
Postman Steps:
- Create new request
- Method:
POST - URL:
http://localhost:8000/predict/lab1-logistic-regression - Headers:
Content-Type: application/json - Body → raw → JSON
- Paste request body
- Click “Send”
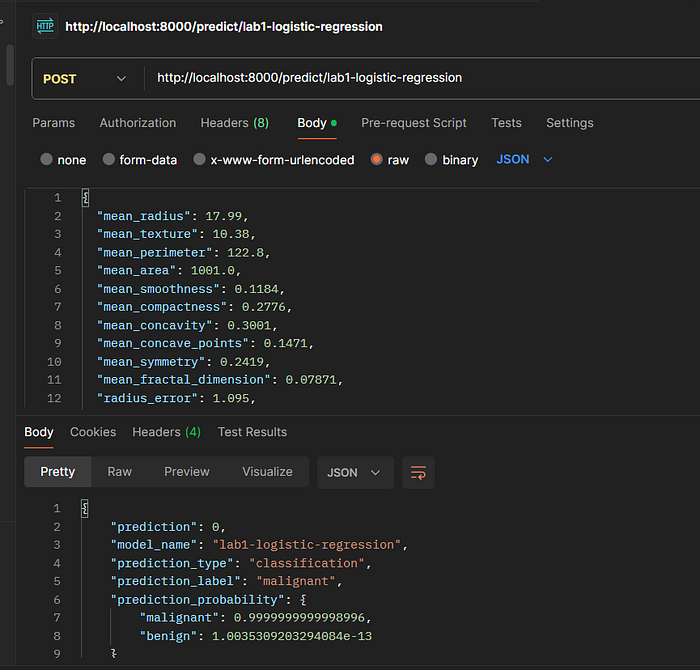
Expected Response:
{
"prediction": 1,
"prediction_label": "benign",
"prediction_probability": {
"malignant": 0.023,
"benign": 0.977
},
"model_name": "lab1-logistic-regression",
"prediction_type": "classification"
}
5.4 Lab 2: Decision Tree Regressor (California Housing)
Endpoint: POST /predict/lab2-decision-tree-regressor
Request Body:
{
"MedInc": 8.3252,
"HouseAge": 41.0,
"AveRooms": 6.98412698,
"AveBedrms": 1.02380952,
"Population": 322.0,
"AveOccup": 2.55555556,
"Latitude": 37.88,
"Longitude": -122.23
}
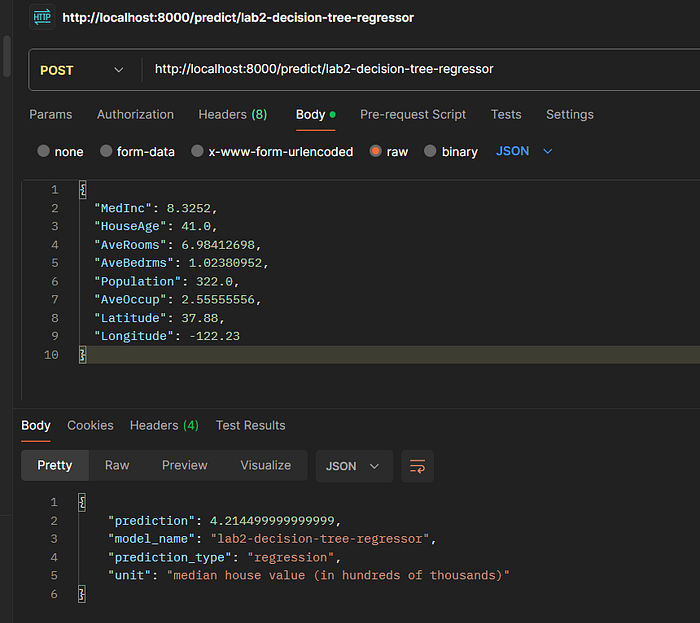
Expected Response:
{
"prediction": 4.526,
"model_name": "lab2-decision-tree-regressor",
"prediction_type": "regression",
"unit": "median house value (in hundreds of thousands)"
}
5.5 Lab 3: K-Means Clustering (Iris)
Endpoint: POST /predict/lab3-kmeans-clustering
Request Body:
{
"sepal_length": 5.1,
"sepal_width": 3.5,
"petal_length": 1.4,
"petal_width": 0.2
}
Expected Response:
{
"prediction": 0,
"cluster_label": "cluster_0",
"model_name": "lab3-kmeans-clustering",
"prediction_type": "clustering"
}
5.6 Lab 4: Random Forest Classifier (Iris)
Endpoint: POST /predict/lab4-random-forest
Request Body:
{
"sepal_length": 5.1,
"sepal_width": 3.5,
"petal_length": 1.4,
"petal_width": 0.2
}
Expected Response:
{
"prediction": 0,
"prediction_label": "setosa",
"prediction_probability": {
"setosa": 0.98,
"versicolor": 0.02,
"virginica": 0.00
},
"model_name": "lab4-random-forest",
"prediction_type": "classification"
}
5.7 Lab 5: Isolation Forest (Anomaly Detection)
Endpoint: POST /predict/lab5-isolation-forest
Request Body: (Same as Lab 1–30 breast cancer features)
Expected Response:
{
"prediction": 1,
"prediction_label": "normal",
"model_name": "lab5-isolation-forest",
"prediction_type": "anomaly_detection"
}
5.8 XGBoost Classifier (Adult Dataset)
Endpoint: POST /predict/xgboost-classifier
Request Body:
{
"Age": 39,
"Workclass": "State-gov",
"Fnlwgt": 77516,
"Education": "Bachelors",
"Education-Num": 13,
"Marital-Status": "Never-married",
"Occupation": "Adm-clerical",
"Relationship": "Not-in-family",
"Race": "White",
"Sex": "Male",
"Capital-Gain": 2174,
"Capital-Loss": 0,
"Hours-per-week": 40,
"Native-Country": "United-States"
}
Expected Response:
{
"prediction": 0,
"prediction_label": "<=50K",
"prediction_probability": {
"<=50K": 0.85,
">50K": 0.15
},
"model_name": "xgboost-classifier",
"prediction_type": "classification"
}
5.9 Linear Regressor (California Housing)
Endpoint: POST /predict/linear-regressor
Request Body: (Same as Lab 2–8 California Housing features)
Expected Response:
{
"prediction": 4.526,
"model_name": "linear-regressor",
"prediction_type": "regression",
"unit": "median house value (in hundreds of thousands)"
}
5.10 Postman Collection Setup
Step 1: Create Collection
- Open Postman
- Click “New” → “Collection”
- Name: “MLflow Prediction API”
Step 2: Create Environment (Optional)
- Click “Environments” → “Create Environment”
- Name: “MLflow Local”
- Add variables:
base_url:http://localhost:8000mlflow_ui:http://localhost:5000
4. Save
Step 3: Add Requests
- Right-click collection → “Add Request”
- Name each request (e.g., “Health Check”, “Lab 1 — Logistic Regression”)
- Configure method, URL, headers, and body for each
Step 4: Use Environment Variables
- In URL field:
{{base_url}}/health - Postman will substitute
{{base_url}}with actual value
5.11 Testing Workflow docker
Complete Testing Sequence:
- Start Services:
docker-compose up -d
2. Train Models:
docker-compose exec mlflow-app python lab1_logistic_regression.py
docker-compose exec mlflow-app python lab2_decision_tree_regressor.py
docker-compose exec mlflow-app python lab3_kmeans_clustering.py
docker-compose exec mlflow-app python lab4_random_forest_classifier.py
docker-compose exec mlflow-app python lab5_isolation_forest.py
docker-compose exec mlflow-app python lab6_wine_quality_autologging.py
docker-compose exec mlflow-app python lab7_wine_quality_manual.py
docker-compose exec mlflow-app python classification.py
docker-compose exec mlflow-app python regressor.py
Note: Lab 6 and Lab 7 require Kaggle API credentials (optional). If not configured, they automatically use UCI ML Repository. See KAGGLE_SETUP.md for setup instructions.
- Verify Health:
- GET
http://localhost:8000/health - Verify
models_loaded: 7
- Test Each Endpoint:
- Use Postman collection or cURL examples
- Verify response format and value
- Error Testing:
- Test with missing fields
- Test with invalid data types
- Test with out-of-range values
5.12 Common Errors and Solutions
503 Service Unavailable:
- Cause: Model not loaded
- Solution: Train the model first, then check
/healthendpoint
422 Validation Error:
- Cause: Invalid request body format
- Solution: Check JSON syntax, required fields, and data types
500 Internal Server Error:
- Cause: Model prediction failed
- Solution: Check API logs:
docker-compose logs mlflow-api
Connection Refused:
- Cause: Services not running
- Solution: Start services:
docker-compose up -d
6. Architecture Diagrams
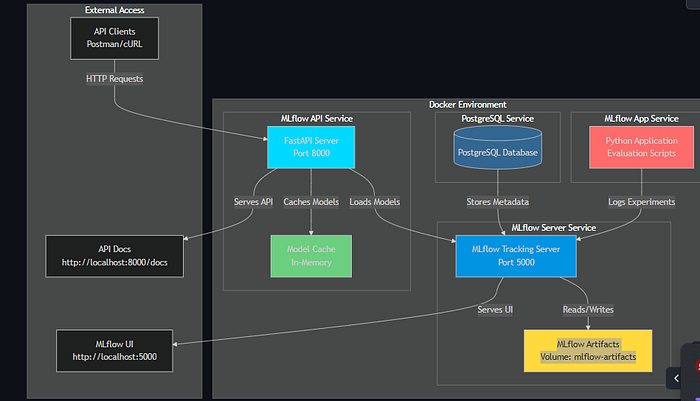
6.1 System Component Diagram
6.2 Data Flow Sequence Diagram
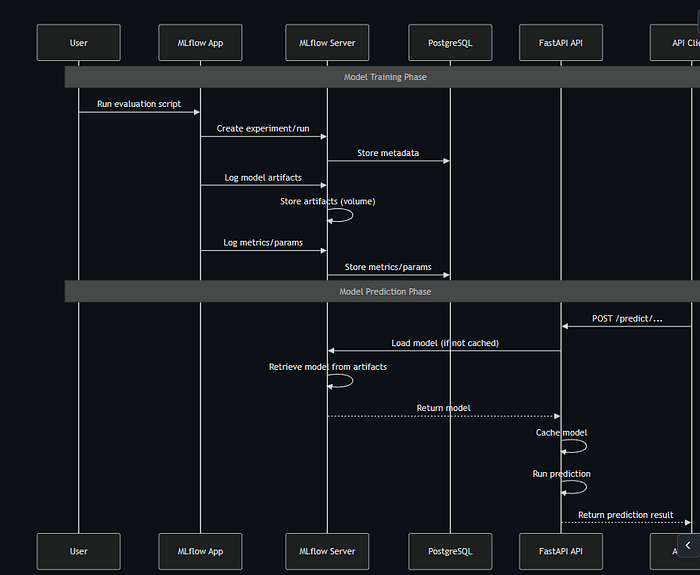
Request Processing Flow
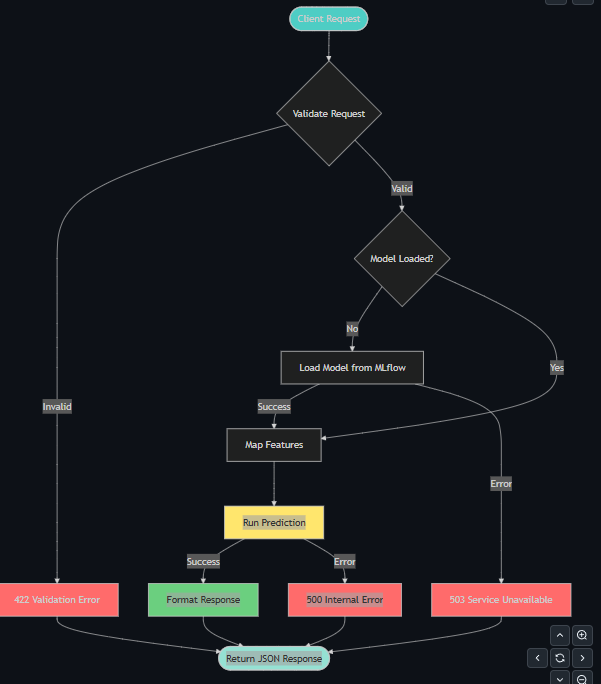
Docker Service Dependencies
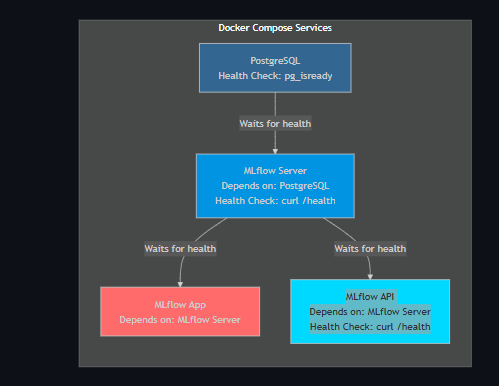
MLflow Experiment Tracking Flow
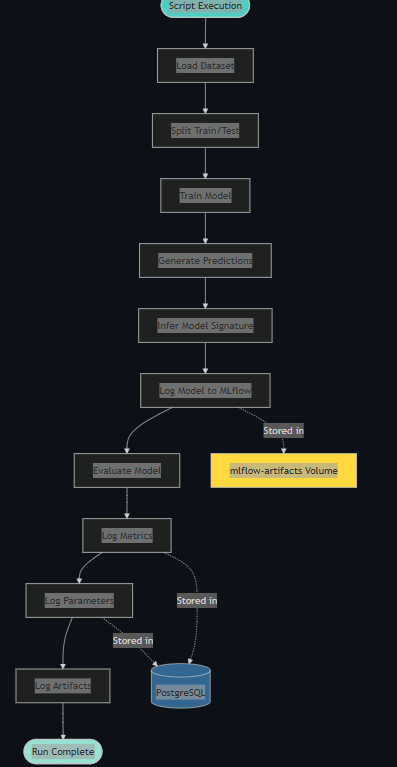
7. Conclusion
This comprehensive guide covers:
- Theoretical Foundations: Deep dive into all 7 machine learning algorithms
- Dataset Understanding: Complete feature descriptions and target explanations (including Wine Quality dataset)
- Code Walkthrough: Step-by-step explanation of all evaluation and prediction services
- Docker Deployment: Complete Docker setup with PostgreSQL backend
- API Testing: Detailed Postman testing guide with all endpoints
- Architecture Visualization: Mermaid diagrams for system design and data flow
- MLflow Logging: Autologging and manual logging demonstrations (Lab 6 & Lab 7)
- Dataset Integration: Kaggle API integration with automatic fallback to UCI ML Repository
The MLflow Evaluation Lab provides a production-ready MLOps workflow with:
- Scalable Architecture: PostgreSQL backend for large-scale experiments
- Reliable Deployment: Docker Compose with health checks and dependency management
- Comprehensive Evaluation: 9 different ML models covering classification, regression, clustering, and anomaly detection
- Flexible Logging: Both autologging and manual logging approaches demonstrated
- Dataset Management: Automated dataset download with Kaggle API support
- RESTful API: FastAPI-based prediction service with lazy loading and error handling
- Complete Documentation: This guide serves as a reference for understanding and extending the system
Next Steps:
- Experiment with different hyperparameters
- Add custom evaluation metrics
- Integrate with CI/CD pipelines
- Deploy to cloud platforms (AWS, GCP, Azure)
- Add model versioning and staging workflows
Github link: https://github.com/faizulkhan56/mlflow_evaluation_lab
Join thousands of data leaders on the AI newsletter. Join over 80,000 subscribers and keep up to date with the latest developments in AI. From research to projects and ideas. If you are building an AI startup, an AI-related product, or a service, we invite you to consider becoming a sponsor.
Published via Towards AI
Towards AI Academy
We Build Enterprise-Grade AI. We'll Teach You to Master It Too.
15 engineers. 100,000+ students. Towards AI Academy teaches what actually survives production.
Start free — no commitment:
→ 6-Day Agentic AI Engineering Email Guide — one practical lesson per day
→ Agents Architecture Cheatsheet — 3 years of architecture decisions in 6 pages
Our courses:
→ AI Engineering Certification — 90+ lessons from project selection to deployed product. The most comprehensive practical LLM course out there.
→ Agent Engineering Course — Hands on with production agent architectures, memory, routing, and eval frameworks — built from real enterprise engagements.
→ AI for Work — Understand, evaluate, and apply AI for complex work tasks.
Note: Article content contains the views of the contributing authors and not Towards AI.